A multi-site genetic study, the Genetic Architecture of Autism (GALA) consortium, has identified shared deleterious coding variations linked to autism spectrum disorder (ASD) across individuals of diverse ancestries, particularly within Latin American populations. This research, published this week, underscores the importance of inclusive genomic studies to understand the complex genetic basis of ASD and improve diagnostic accuracy globally.
Understanding the Genetic Landscape of Autism
Autism Spectrum Disorder is a complex neurodevelopmental condition characterized by challenges with social communication and interaction, as well as restricted, repetitive patterns of behavior, interests, or activities. While environmental factors are believed to play a role, genetic predisposition is a significant contributor, with estimates suggesting that heritability accounts for 80% of the risk. However, the genetic architecture of ASD remains largely unresolved, and most identified genetic variants explain only a small fraction of the overall heritability. This new research from GALA, involving over 14,000 individuals, sheds light on the shared genetic underpinnings of ASD, irrespective of ancestral background.
In Plain English: The Clinical Takeaway
- Shared Genes Matter: This study confirms that some of the same genetic changes contribute to autism in people from different parts of the world, meaning genetic testing can be helpful for diagnosis regardless of ancestry.
- More Inclusive Research is Key: Historically, genetic studies have focused on individuals of European descent. This research highlights the importance of including diverse populations to get a complete picture of the genetic factors involved in autism.
- Personalized Medicine Potential: Identifying these shared genetic variations could eventually lead to more targeted therapies and interventions for individuals with autism.
The GALA Consortium and Ancestry-Specific Findings
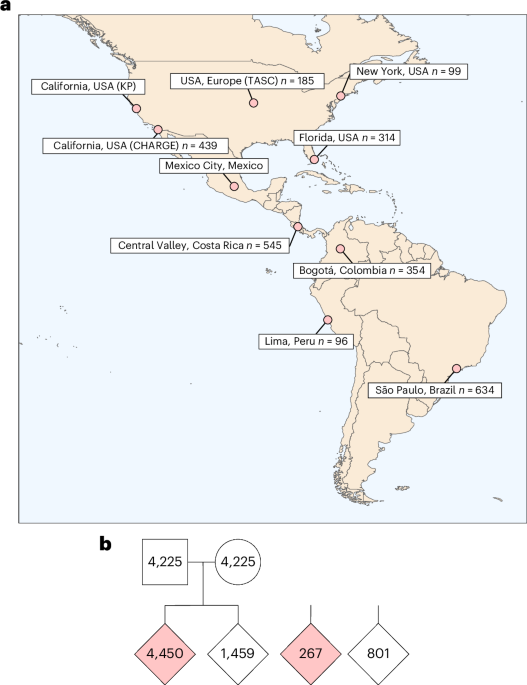
The GALA consortium is a collaborative effort involving research sites across North, Central, and South America. The study meticulously analyzed genetic data from individuals with ASD and their families, focusing on identifying rare coding variations – changes in the DNA sequence that alter the protein produced. A key aspect of this research was its emphasis on individuals of Ancestry Informative Markers (AIM) from the Americas (AMR), a population historically underrepresented in genetic studies. Researchers utilized a sophisticated approach to determine ancestry, combining genetic data with principal component analysis and a random forest classifier trained on reference populations.

The study identified several genes with recurrent deleterious variants in individuals with ASD, including SYNGAP1, GRIN2B, and CHD8. Importantly, the frequency of these variants was comparable across different AMR subgroups, suggesting a shared genetic vulnerability. The researchers similarly investigated copy number variations (CNVs) – deletions or duplications of large segments of DNA – and found that these events also contribute to ASD risk across ancestries.
Clinical Genetics and ACMG Interpretation
Beyond identifying specific genes, the researchers also applied the American College of Medical Genetics and Genomics (ACMG) guidelines to interpret the clinical significance of the identified variants. This involved assessing the evidence for pathogenicity based on factors such as the type of mutation, its frequency in the population, and its predicted impact on protein function. While the ACMG guidelines are typically applied to individual cases, the researchers used a modified approach to analyze the collective data, focusing on genes with a strong association with autism and related neurodevelopmental disorders. They utilized tools like VarSome and Neptune to aid in variant annotation and classification.
The use of Neptune, a genomic medicine environment, revealed that the frequency of potentially pathogenic/likely pathogenic (P/LP) variants was similar between AMR and non-AMR participants, although Neptune had a slightly reduced ability to classify variants in the AMR group. This highlights the need for continued refinement of genomic tools to ensure equitable performance across diverse populations.
Funding and Bias Transparency
This research was supported by grants from the National Institutes of Health (NIH), Autism Speaks, and the Simons Foundation. This proves vital to acknowledge that funding sources can potentially influence research priorities and interpretations. However, the GALA consortium has taken steps to mitigate bias by involving investigators from multiple institutions and ensuring transparency in data analysis and reporting.
“Our findings underscore the importance of diversifying genomic research to ensure that all individuals, regardless of their ancestry, benefit from advances in precision medicine,” states Dr. Matthew State, a lead investigator on the GALA consortium. “By including historically underrepresented populations, You can gain a more complete understanding of the genetic basis of autism and develop more effective diagnostic and therapeutic strategies.”
Geographical Impact and Healthcare Systems
The implications of this research extend beyond the laboratory and into clinical practice. In the United States, the findings could influence the implementation of genetic testing guidelines for ASD, potentially leading to earlier and more accurate diagnoses. The Food and Drug Administration (FDA) is currently evaluating several novel therapies for ASD, and the identification of shared genetic targets could accelerate the development of personalized treatments. Similarly, in Europe, the European Medicines Agency (EMA) is actively involved in assessing the safety and efficacy of new ASD therapies, and this research could inform their regulatory decisions. In Latin America, where access to genetic testing may be limited, the findings emphasize the need for increased investment in genomic infrastructure and training of healthcare professionals.
Contraindications & When to Consult a Doctor
Genetic testing for ASD is not a standalone diagnostic tool. It should always be interpreted in the context of a comprehensive clinical evaluation by a qualified healthcare professional. Individuals with a family history of ASD or developmental delays should consult with a genetic counselor or pediatrician to discuss the potential benefits and limitations of genetic testing. It’s crucial to understand that a positive genetic test result does not necessarily signify that an individual will develop ASD, and a negative result does not rule out the possibility. Individuals experiencing anxiety or distress related to genetic testing results should seek support from a mental health professional.
Data Summary: Variant Classification
| Variant Type | AMR (N=4450) | Non-AMR (N=13030) |
|---|---|---|
| Pathogenic/Likely Pathogenic (P/LP) | 136 (1.60%) | 344 (1.66%) |
| Variants Classified by Neptune | 8501 (69.9%) | 20750 (73.4%) |
| Total Variants Analyzed | 12162 | 28262 |
Future Directions and Longitudinal Studies
The GALA consortium is continuing to expand its research efforts, with plans to recruit additional participants and investigate the interplay between genetic and environmental factors in ASD. Longitudinal studies, which follow individuals with ASD over time, are crucial for understanding the long-term trajectory of the disorder and identifying potential targets for intervention. Research is needed to develop more effective therapies for ASD, addressing the core symptoms of social communication deficits and repetitive behaviors.
“This represents just the beginning,” says Dr. Lisa Croen, a leading autism researcher at Kaiser Permanente. “By continuing to collaborate and share data, we can unlock the mysteries of autism and improve the lives of individuals and families affected by this complex condition.”
References
- Fu, J. M., et al. (2022). Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nature Genetics, 54(11), 1320–1331. https://doi.org/10.1038/s41588-022-01211-9
- Satterstrom, F. K., et al. (2020). Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell, 180(5), 568–584.e19. https://doi.org/10.1016/j.cell.2020.01.032
- Venner, E., et al. (2024). The frequency of pathogenic variation in the All of Us cohort reveals ancestry-driven disparities. Communications Biology, 7(1), 174. https://doi.org/10.1038/s42003-024-01641-x
- Autism Speaks. (n.d.). What is Autism? https://www.autismspeaks.org/what-is-autism
- Centers for Disease Control and Prevention (CDC). (2023). Autism Spectrum Disorder (ASD). https://www.cdc.gov/ncbddd/autism/index.html