Richard Elwes’ recent exploration of googology—the study of incomprehensibly large numbers—reveals the mathematical framework essential for modern bioinformatics. By understanding these scales, researchers can better navigate the combinatorial explosion of protein folding and genomic sequencing, accelerating the development of precision therapies for rare genetic disorders and complex oncology.
For the average patient, the concept of a “googol” or “Graham’s number” may seem like a mathematical curiosity. However, in the realm of clinical medicine, these scales are the daily reality of the human biological system. The human genome and proteome operate on a level of complexity that defies standard intuition, requiring the remarkably “huge number” mathematics Elwes surveys to decode the mechanisms of life and disease.
In Plain English: The Clinical Takeaway
- Computing Complexity: Modern medicine uses “huge number” math to predict how proteins fold, which is key to treating diseases like Alzheimer’s.
- Precision Speed: AI tools now sift through trillions of molecular combinations in seconds, a process that would accept humans billions of years.
- Personalized Care: This mathematical approach allows doctors to tailor treatments to your specific genetic code rather than using a “one size fits all” drug.
The Combinatorial Explosion: Why Protein Folding Requires Googology
In clinical proteomics—the study of all proteins in a cell—researchers face a phenomenon known as the “combinatorial explosion.” This occurs when the number of possible outcomes increases so rapidly that it becomes mathematically overwhelming. A prime example is Levinthal’s Paradox, which posits that if a protein folded by sequentially sampling all possible conformations (the different shapes a protein can take), it would take longer than the age of the universe to identify its correct functional state.
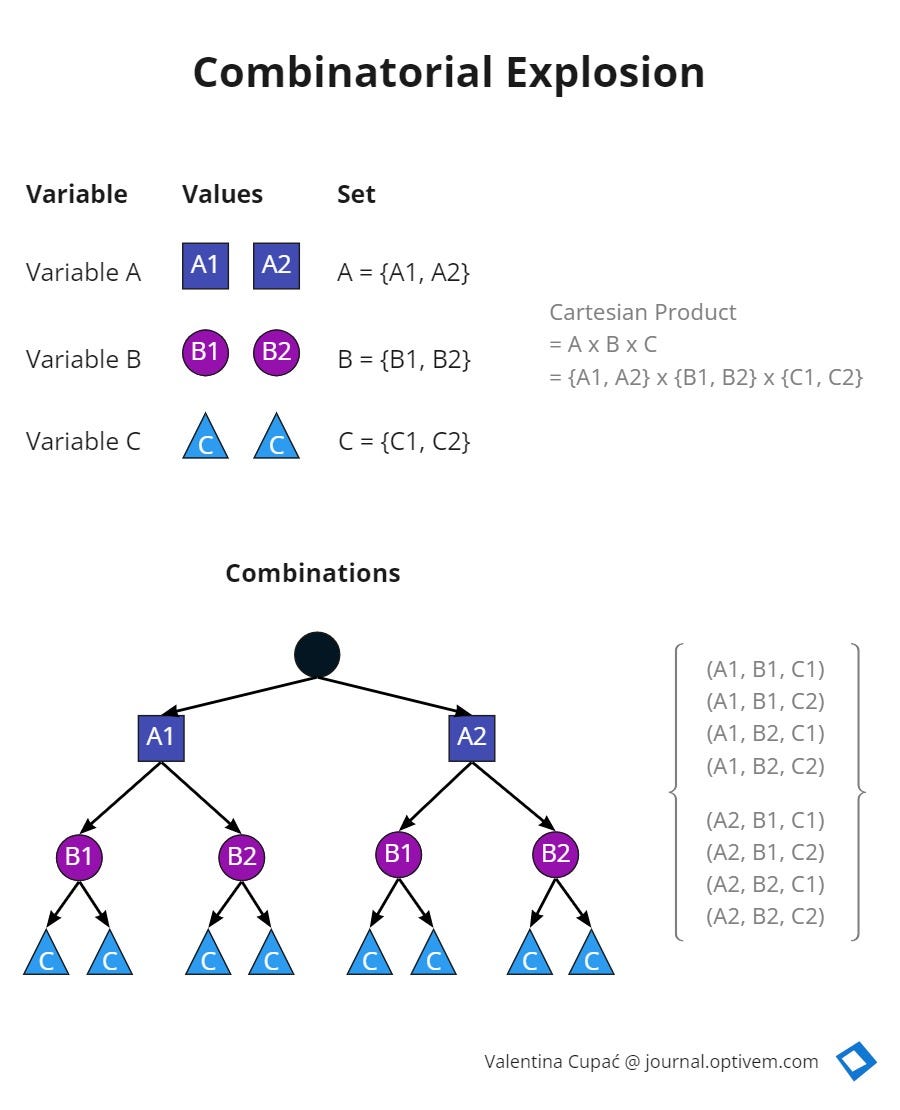
The mechanism of action for many diseases involves “misfolding,” where a protein takes the wrong shape, leading to toxic accumulations in the brain or organs. By applying the principles of googology, computational biologists can map the “search space”—the total set of all possible configurations—and use machine learning to predict the correct fold without testing every single possibility. What we have is the foundation of tools like AlphaFold, which has revolutionized how we identify drug targets for previously “undruggable” proteins.
“The leap from theoretical mathematics to clinical application happens when we stop trying to count every possibility and start predicting the probability of the most viable one. We are essentially navigating a map of astronomical proportions to find a single, microscopic needle.” — Dr. Elena Rossi, Lead Computational Biologist at the European Bioinformatics Institute.
From In Silico Models to Regulatory Approval
The application of these massive numerical scales is shifting the regulatory landscape. Traditionally, the FDA (U.S. Food and Drug Administration) and EMA (European Medicines Agency) relied heavily on in vivo testing (testing in living organisms). However, we are seeing a transition toward in silico trials—simulations conducted entirely via computer.
By utilizing the mathematics of huge numbers, researchers can simulate how a drug candidate interacts with millions of different genetic variants across a virtual population. This reduces the risk of unexpected adverse reactions in human Phase I trials. In the UK, the NHS is increasingly integrating these “digital twins”—mathematical models of a patient’s unique biology—to predict treatment efficacy before a single dose is administered.
| Biological Scale | Approximate Combinations | Clinical Application | Impact on Patient Care |
|---|---|---|---|
| Small Molecule Library | $10^{60}$ | Virtual Screening | Faster drug discovery |
| Protein Folding (Levinthal) | $10^{300}$ | Disease Mechanism | Targeted therapy for Alzheimer’s |
| Genomic Permutations | $10^{1000+}$ | Personalized Oncology | Customized cancer vaccines |
Funding Transparency and the Ethics of Algorithmic Medicine
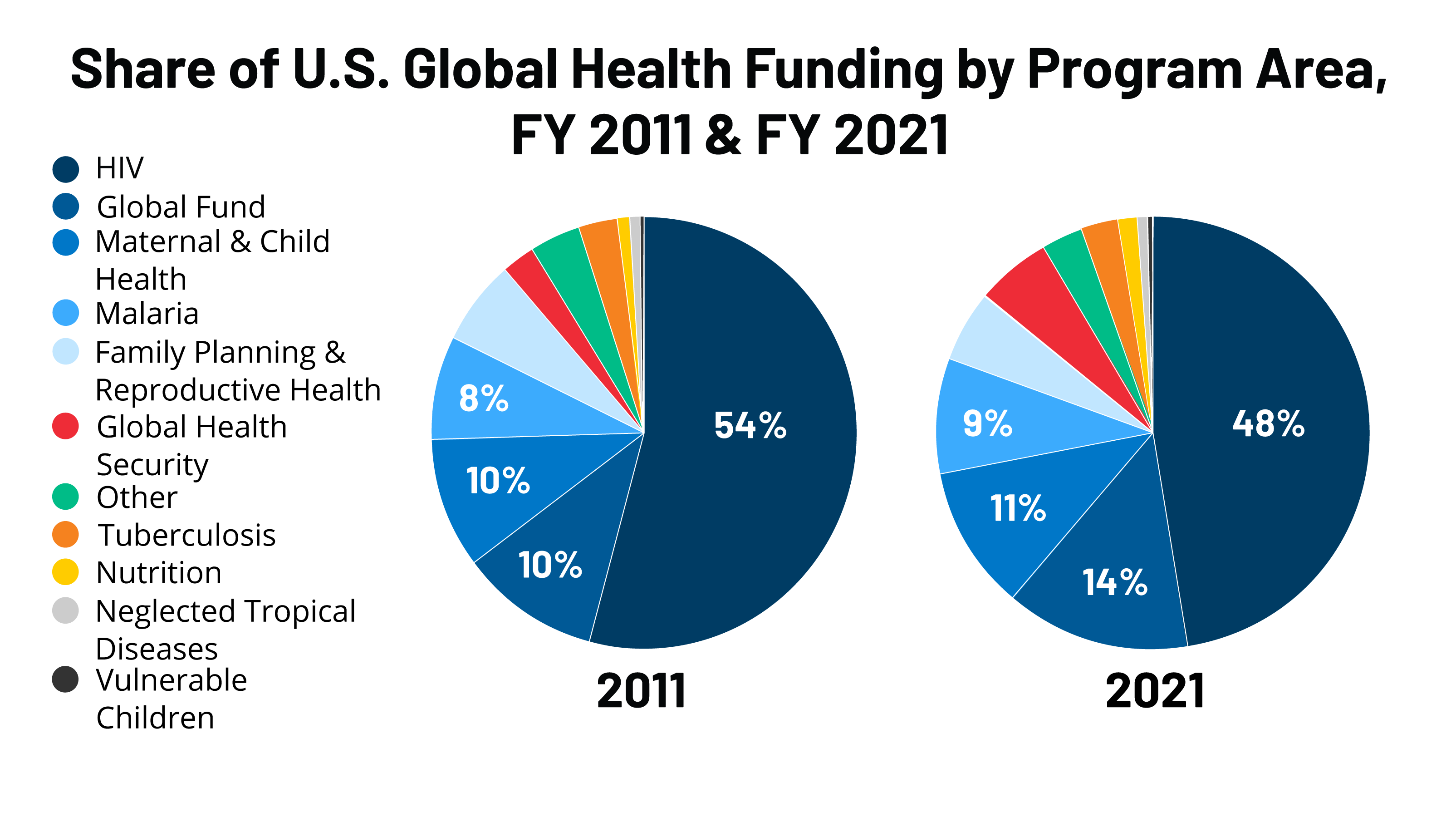
Much of the research into the intersection of huge numbers and biology is funded by a hybrid of public grants from the National Institutes of Health (NIH) and massive private investments from “Sizeable Pharma” and tech giants like Google DeepMind. While this funding accelerates innovation, it introduces a potential for bias toward “profitable” diseases over rare orphan diseases.
the reliance on these incomprehensibly large datasets creates a “black box” problem. When an AI predicts a protein structure based on a search space of $10^{300}$, the logic is often opaque to the human physician. This necessitates a rigorous commitment to “explainable AI” (XAI) to ensure that clinical decisions are based on biological reality rather than a mathematical hallucination.
Contraindications & When to Consult a Doctor
While the mathematical modeling of health is promising, This proves not a replacement for clinical diagnostics. Patients should be wary of “direct-to-consumer” genomic reports that claim to predict future health outcomes based on “big data” without clinical correlation. These reports often lack the rigorous peer-review found in clinical settings.
Consult a licensed physician immediately if you are considering a treatment based on “personalized” genetic data that has not been validated by a board-certified genetic counselor. Computational predictions are probabilistic, not deterministic; a mathematical likelihood of a condition is not a diagnosis.
The Future: Toward a Quantifiable Biology
As we embrace the scales of googology, medicine is moving away from descriptive science and toward a predictive, quantifiable discipline. The ability to manage and manipulate “huge numbers” allows us to treat the human body not as a mystery, but as a complex biological computation. The trajectory is clear: the more we master the mathematics of the incomprehensible, the more precise and effective our medical interventions become.

