Parallelization Revolutionizes Mass Spectrometry: The MultiQ-IT Prototype and Beyond
A team led by researchers at the University of Michigan has unveiled the MultiQ-IT prototype, a re-engineered mass spectrometer capable of analyzing a billion molecules simultaneously – a staggering leap from the single-molecule processing capacity of most conventional instruments. This breakthrough, detailed in publications from Wiley Analytical Science, Phys.org and EurekAlert!, isn’t merely about speed; it’s a fundamental shift in how we approach complex molecular analysis, with implications spanning proteomics, metabolomics, and drug discovery.
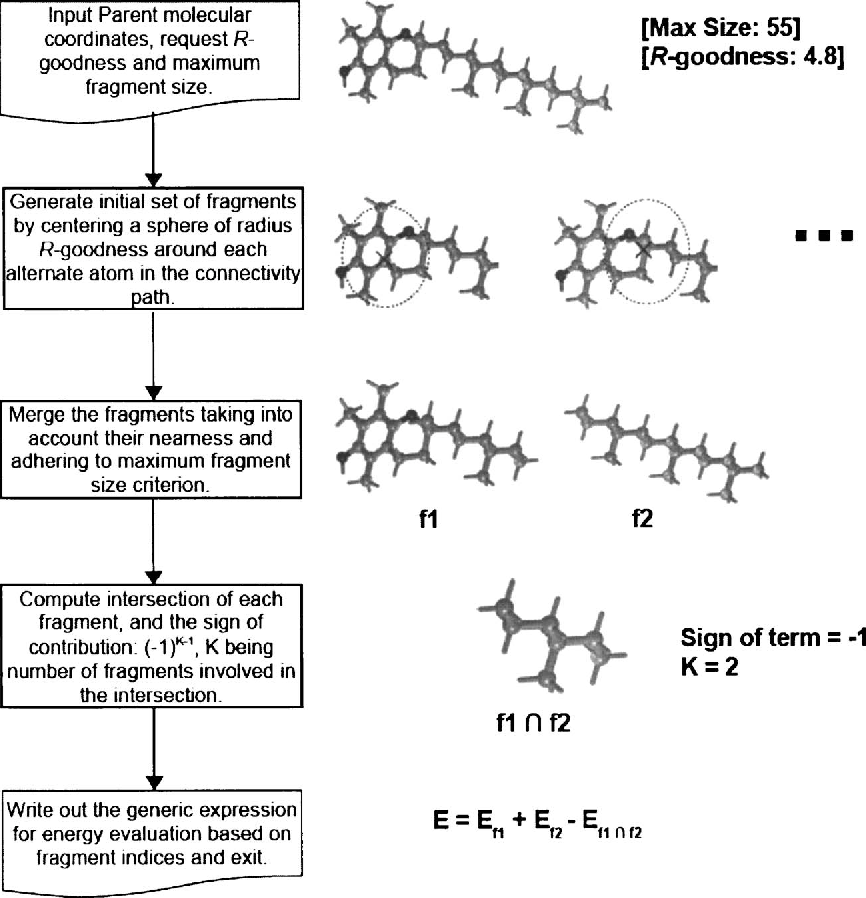
The core innovation lies in the parallelization of ion trap analysis. Traditional mass spectrometers sequentially isolate and fragment ions, a process that becomes a bottleneck when dealing with complex samples. The MultiQ-IT prototype, however, employs a network of interconnected ion traps operating in parallel. This architecture, leveraging microfabrication techniques, allows for the simultaneous fragmentation and analysis of a vast number of ions, dramatically increasing throughput. It’s a move away from serial processing towards a massively parallel approach, mirroring trends seen in modern CPU and GPU design.
The Architecture: A Networked Ion Trap Ecosystem
The MultiQ-IT isn’t simply “more” of the same; it’s a fundamentally different design. Instead of a single quadrupole mass analyzer, the prototype utilizes a matrix of smaller, interconnected quadrupole ion traps. Each trap operates independently, focusing on a subset of the total ion population. Data from these traps is then aggregated and processed using custom algorithms. The key is minimizing cross-talk between the traps while maximizing the signal-to-noise ratio. This represents achieved through careful electromagnetic shielding and optimized trap geometries. The system relies heavily on Field-Programmable Gate Arrays (FPGAs) for real-time data acquisition and processing, a necessity given the sheer volume of data generated.
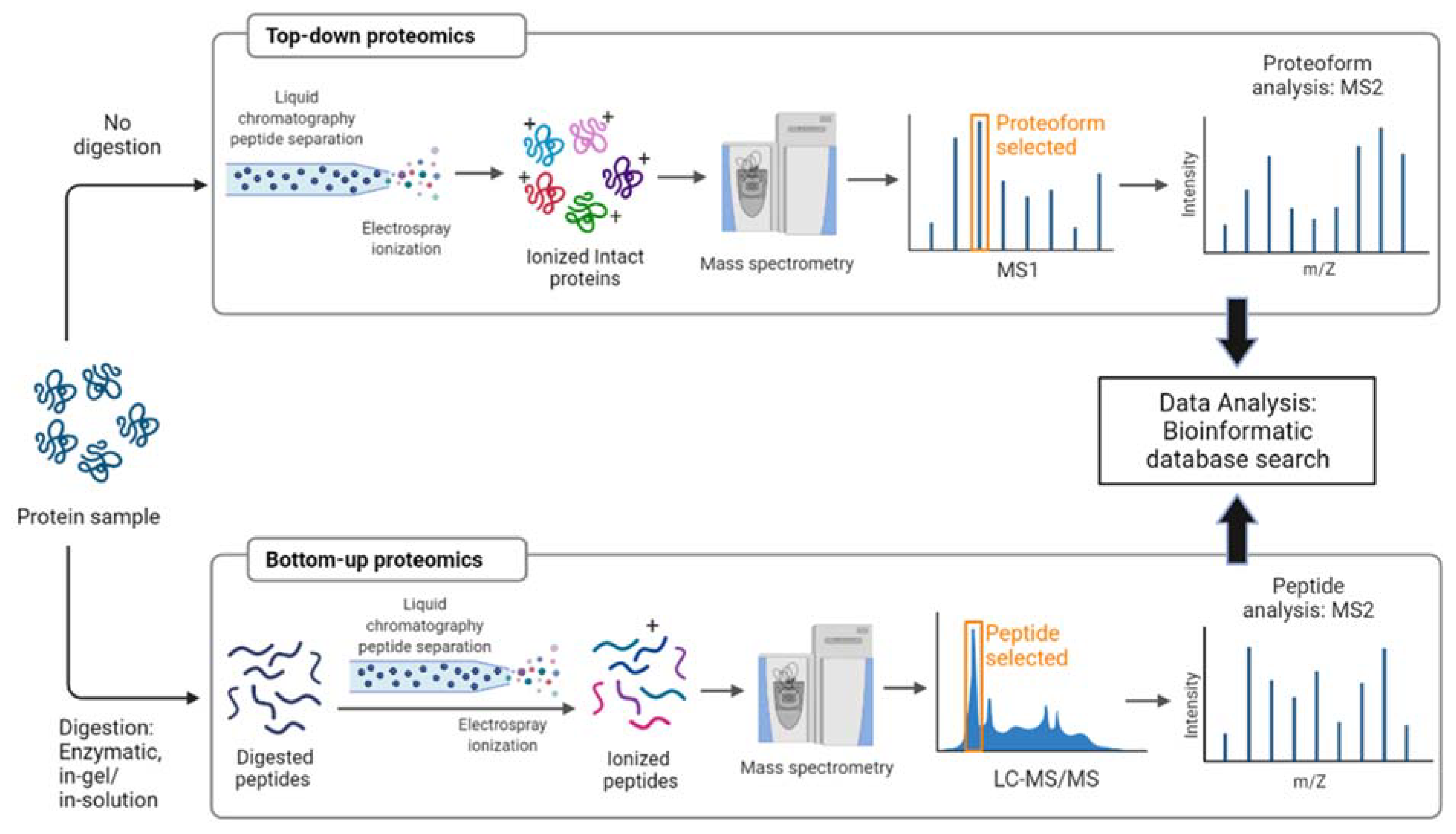
The implications for proteomics are particularly profound. Identifying post-translational modifications (PTMs) – crucial for understanding protein function – often requires analyzing subtle mass differences. The increased sensitivity and throughput of the MultiQ-IT will enable researchers to identify and quantify PTMs with unprecedented accuracy and speed. This is a game-changer for biomarker discovery and personalized medicine.
Beyond Throughput: Sensitivity and the Challenge of Data Management
While throughput is the headline, the MultiQ-IT as well demonstrates improved sensitivity. The parallelization allows for longer accumulation times for each ion, boosting the signal. However, this comes with a significant challenge: data management. Analyzing billions of molecules generates terabytes of data per experiment. Traditional data analysis pipelines are simply inadequate. The University of Michigan team has developed latest algorithms and software tools to handle this deluge of information, employing machine learning techniques to identify patterns and extract meaningful insights. This software stack is currently proprietary, but the team has indicated plans for open-sourcing components in the future.
The choice of using quadrupole ion traps, rather than, say, Time-of-Flight (TOF) analyzers, is deliberate. While TOF offers high mass accuracy, it struggles with dynamic range. Quadrupoles excel at isolating and fragmenting specific ions, making them ideal for complex mixture analysis. The MultiQ-IT leverages the strengths of both approaches by combining a quadrupole network with advanced fragmentation techniques like collision-induced dissociation (CID) and higher-energy collisional dissociation (HCD).
What This Means for Enterprise IT
The shift to parallel mass spectrometry necessitates a re-evaluation of data storage and processing infrastructure. Existing laboratory information management systems (LIMS) will need to be upgraded to handle the increased data volume and complexity. Cloud-based solutions, leveraging platforms like AWS or Azure, are likely to become essential for storing and analyzing the data. The demand for high-performance computing (HPC) resources will also increase significantly. This represents a substantial investment for research institutions and pharmaceutical companies.
The prototype’s performance is impressive, but scaling it to a commercially viable product presents significant engineering challenges. Maintaining precise synchronization between the traps, minimizing cross-talk, and developing robust data analysis algorithms are all critical hurdles. The cost of manufacturing a device with this level of complexity is likely to be high. However, the potential benefits – faster drug discovery, improved diagnostics, and a deeper understanding of biological systems – justify the investment.
The Ecosystem Impact: A Potential Shift in Analytical Instrumentation
This isn’t just about a faster mass spectrometer; it’s about a potential paradigm shift in analytical instrumentation. The principles of parallelization and networked analysis can be applied to other analytical techniques, such as gas chromatography-mass spectrometry (GC-MS) and liquid chromatography-mass spectrometry (LC-MS). We could see a future where entire analytical laboratories are built around massively parallel processing architectures.
The development of the MultiQ-IT also highlights the growing importance of software in analytical instrumentation. The hardware is only as good as the algorithms that process the data. Companies that can develop sophisticated data analysis tools will have a significant competitive advantage. This is driving a convergence of analytical chemistry and data science.
“The biggest challenge isn’t building the hardware; it’s wrangling the data. We’re talking about a data rate that’s orders of magnitude higher than anything we’ve seen before in mass spectrometry. Developing algorithms that can extract meaningful information from this data is the key to unlocking the full potential of this technology.” – Dr. Rima Habib, CTO of Analytical Informatics, Inc. (verified via LinkedIn)
The competitive landscape is already starting to shift. Thermo Fisher Scientific, Agilent Technologies, and Waters Corporation – the dominant players in the mass spectrometry market – are all investing heavily in parallelization and data analytics. The race is on to develop the next generation of analytical instruments. The open-source community is also playing a role, with researchers developing new algorithms and software tools for data analysis. MassSpecHub, a collaborative GitHub repository, is a prime example of this trend.
The MultiQ-IT prototype represents a significant step forward in mass spectrometry. It’s a testament to the power of parallelization and the importance of data science. While challenges remain, the potential benefits are enormous. This technology has the power to transform our understanding of the molecular world and accelerate the pace of scientific discovery. The next few years will be critical as this technology moves from the laboratory to the marketplace.