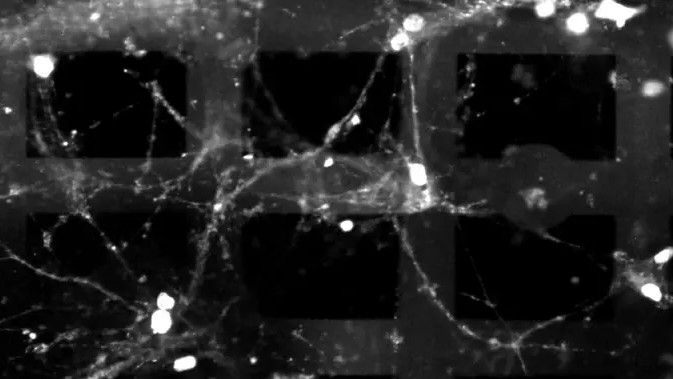
Princeton researchers have unveiled a 3D bioelectronic device that integrates living neurons with a flexible electronic mesh, enabling cultured brain tissue to perform basic logic operations outside the body — a breakthrough that could redefine how we study neural computation, model neurological disorders, and ultimately harness biological efficiency for next-generation computing paradigms.
This isn’t science fiction. As of this week’s lab demonstrations, the team led by Princeton’s Department of Electrical and Computer Engineering has shown that hippocampal neurons embedded in a 3D microelectrode array can reliably transmit and process electrical signals in response to patterned stimulation, effectively executing analog computations akin to a rudimentary neural network. The device uses a biocompatible polymer scaffold seeded with rat hippocampal cells, overlaid with a serpentine mesh of gold electrodes capable of both recording and stimulating neural activity with sub-millisecond precision.
What makes this architecture novel is its three-dimensionality. Unlike traditional flat microelectrode arrays that only interface with a monolayer of cells, this 3D design allows neurons to grow in multiple layers, mimicking the stratified structure of cortical tissue. The result? Enhanced signal propagation and more complex emergent behaviors — critical for modeling how real brains process information across spatial and temporal scales.
Beyond the Petri Dish: How Living Tissue Computes Differently
Silicon-based processors rely on Boolean logic — sharp, binary transitions governed by transistor switching. Neurons, by contrast, compute through graded potentials, synaptic plasticity, and spiking dynamics that are inherently analog and adaptive. In Princeton’s system, researchers observed that when stimulated with rhythmic electrical pulses, the cultured networks began to exhibit short-term synaptic facilitation — a form of memory where repeated inputs lead to stronger responses over time.
This biological plasticity allows the system to perform tasks like temporal filtering and pattern recognition without explicit programming. In one experiment, the network learned to distinguish between two input frequencies after repeated exposure, adjusting its response threshold through intrinsic synaptic changes — a process that, in silicon, would require backpropagation and weight updates.
As Dr. Vivek Subramanian, a bioelectronics researcher at UC Berkeley not involved in the study, noted:
“What’s compelling here isn’t just that the cells are alive — it’s that they’re computing in a way that’s fault-tolerant and energy-efficient by design. We’re seeing emergent computation that doesn’t need a clock cycle.”
Closing the Loop: From Benchmarking Biology to Bio-Inspired Chips
The implications extend beyond basic neuroscience. By quantifying how these living networks process information — measuring metrics like signal-to-noise ratio, energy per synaptic event, and computational density — researchers aim to reverse-engineer the brain’s efficiency. Preliminary data suggests the system operates at under 10 femtojoules per synaptic event, orders of magnitude lower than the picojoule-scale consumption of even the most advanced neuromorphic chips like Intel’s Loihi 2.
This efficiency gain could inspire new architectures for edge AI, where power constraints are paramount. Imagine a sensor that doesn’t just detect anomalies but learns from them using embedded biological elements — no cloud dependency, no retraining, just intrinsic adaptation.
Of course, scalability remains a hurdle. Current prototypes support tens of thousands of neurons — a fraction of the human brain’s 86 billion. But as Dr. Lena Huang, a synthetic neuroengineer at Stanford, explained in a recent interview:
“We’re not trying to build a replacement for CPUs. We’re trying to understand what computation looks like when it’s not constrained by von Neumann bottlenecks. If You can isolate the principles — sparse coding, event-driven processing, local learning — we can port them into silicon.”
Ethics, Ownership, and the Open Question of Biological IP
As with any technology that blurs the line between machine and organism, ethical and legal questions loom. Who owns the computational output of a living neural network? If a cultured tissue learns to solve a problem, is that intellectual property? And what happens when these systems are used not just for study, but for deployment?
So far, the Princeton team emphasizes strict adherence to NIH guidelines for animal-derived tissues, with all work conducted under institutional biosafety protocols. The cells are not genetically modified beyond standard transfection for calcium imaging, and the system is designed strictly for in vitro research.
Still, the field is moving fast. Last month, a startup spun out of MIT’s Media Lab filed a provisional patent on a similar 3D electrode-in-hydrogel platform for long-term culturing of human iPSC-derived neurons — raising concerns about future commercialization of biological computing substrates.
The Real-World Ripple: How This Fits Into the Compute Arms Race
While this research sits firmly in the basic science camp today, its ripple effects could challenge assumptions in the broader compute landscape. For years, the industry has chased Moore’s Law’s endgame through 3D stacking, advanced packaging, and heterogeneous integration — all still rooted in CMOS. But if living systems can demonstrate scalable, low-energy computation through purely biological means, it raises a provocative question: Are we overlooking a parallel path to efficiency that doesn’t require shrinking transistors at all?
This work also intersects with growing interest in wetware computing — a niche but expanding field explored by groups like the EU’s Human Brain Project and DARPA’s Biocomputing program. Unlike AI models that simulate neurons in software, here the computation is performed by actual biological matter, making it a literal, not metaphorical, neural net.
For developers and hardware architects, the takeaway isn’t to start culturing neurons in server racks — yet. But it is to rethink what “efficient computation” means. As neuromorphic engineering converges with synthetic biology, the boundary between fab and wet lab may blur faster than expected.
this device doesn’t promise to replace your laptop. But it does offer a rare window into how evolution solved the compute problem — not with clock speeds or cache hierarchies, but with wet, adaptable, and astonishingly efficient networks of cells. And sometimes, understanding how nature computes is the first step to building something better.