
Former Central Lombok Regent Jailed Over Fraud Case
The transition from the mahogany desks of regional power to the sterile confines of a prison cell often happens in a heartbeat, punctuated by the cold, metallic click of handcuffs. ... Read More
Saturday Edition
Stay updated with Archyde – your source for breaking news, global headlines, economy, entertainment, health, technology, and sports. Fresh stories daily.

The transition from the mahogany desks of regional power to the sterile confines of a prison cell often happens in a heartbeat, punctuated by the cold, metallic click of handcuffs. ... Read More
Continuous Coverage

Severe heatwaves gripping the US Southwest in early May 2026 are straining power grids and water reserves, threatening…
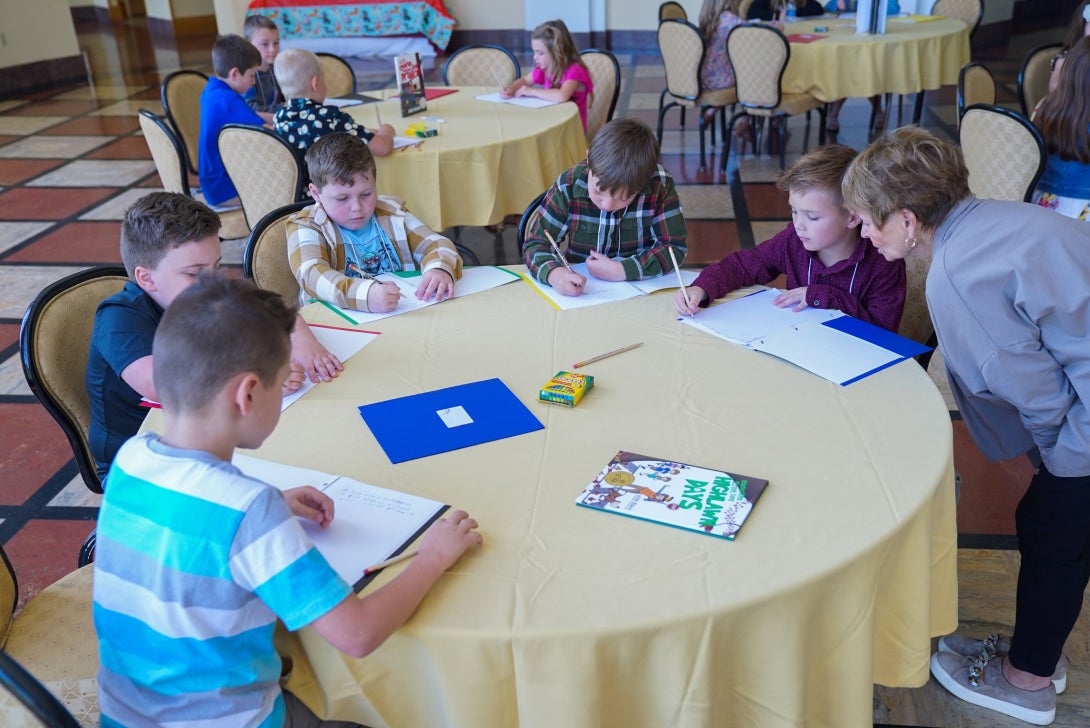

More than West Virginia Department of Education (WVDE) partners and 250 aspiring young authors recently converged in Charleston,…

Strasbourg is a city of contradictions, where half-timbered houses from the 16th century lean precariously against the sleek,…

La p’tite débrouille, a critically acclaimed comedy centered on wartime tolerance, arrives at the Avignon Festival this May…

Female leadership in Italian SMEs, highlighted by emerging hubs like Pescara, is driving regional economic resilience. By integrating…
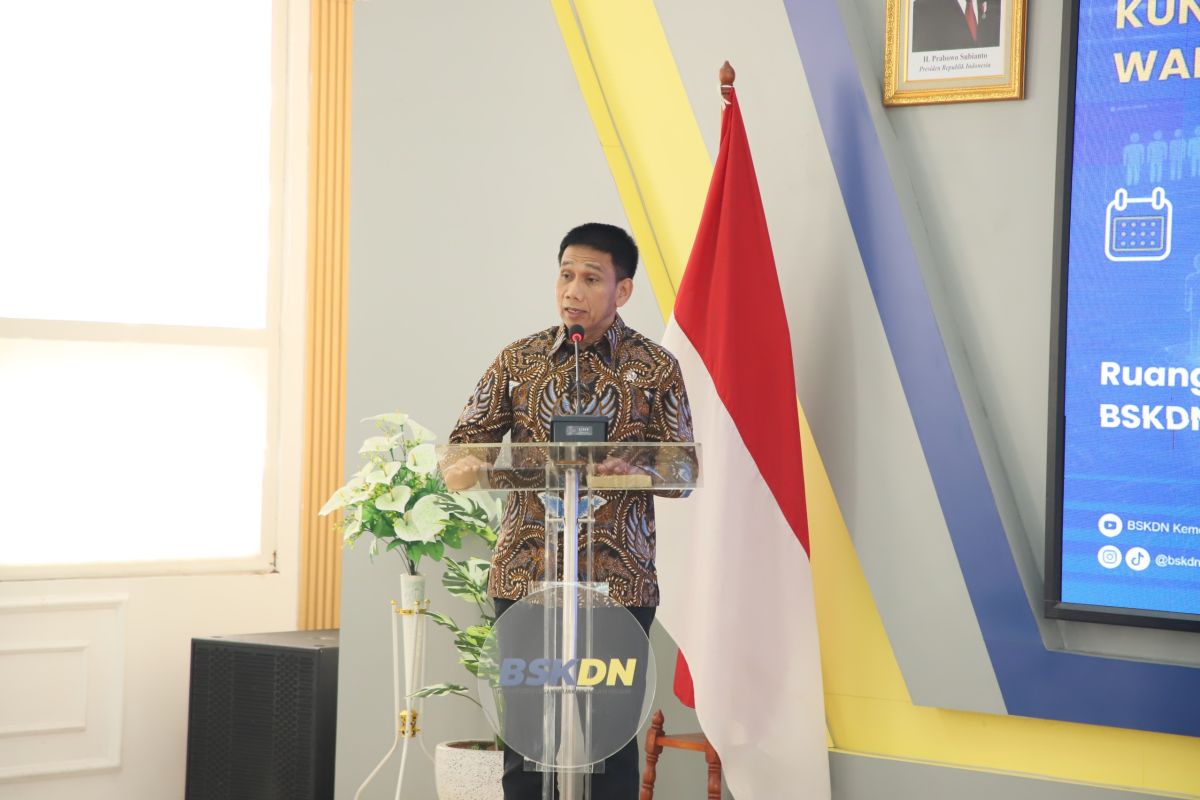

The Indonesian government is launching DESLab, a digital election “sandbox,” to test and refine AI-driven policies and digital…
Global Affairs

Brenda Heist vanished in February 2002 shortly after dropping her children off at school in Pennsylvania. The disappearance…
Markets And Money

Takashi Tezuka, a foundational creative director at Nintendo (TYO: 7974) and key architect of the Mario franchise, is…
Digital Culture

Apple has introduced 3D Spatial Wallpapers in iOS 26, utilizing advanced depth-mapping and NPU-driven parallax effects to create…
Science And Wellbeing
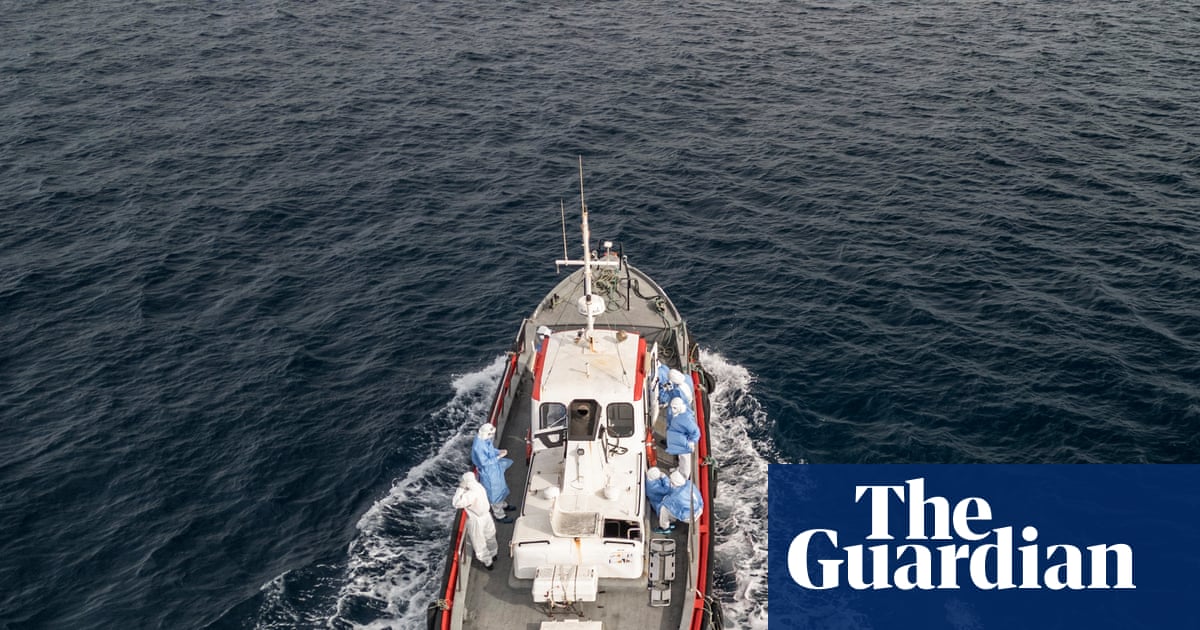

A hantavirus outbreak aboard the cruise ship MV Hondius has highlighted critical vulnerabilities in the US public health…
Screen And Sound

On Friday night, May 8, 2026, US forces struck two Iranian-flagged oil tankers, including the sanctioned vessel Jin…
Fixtures And Form
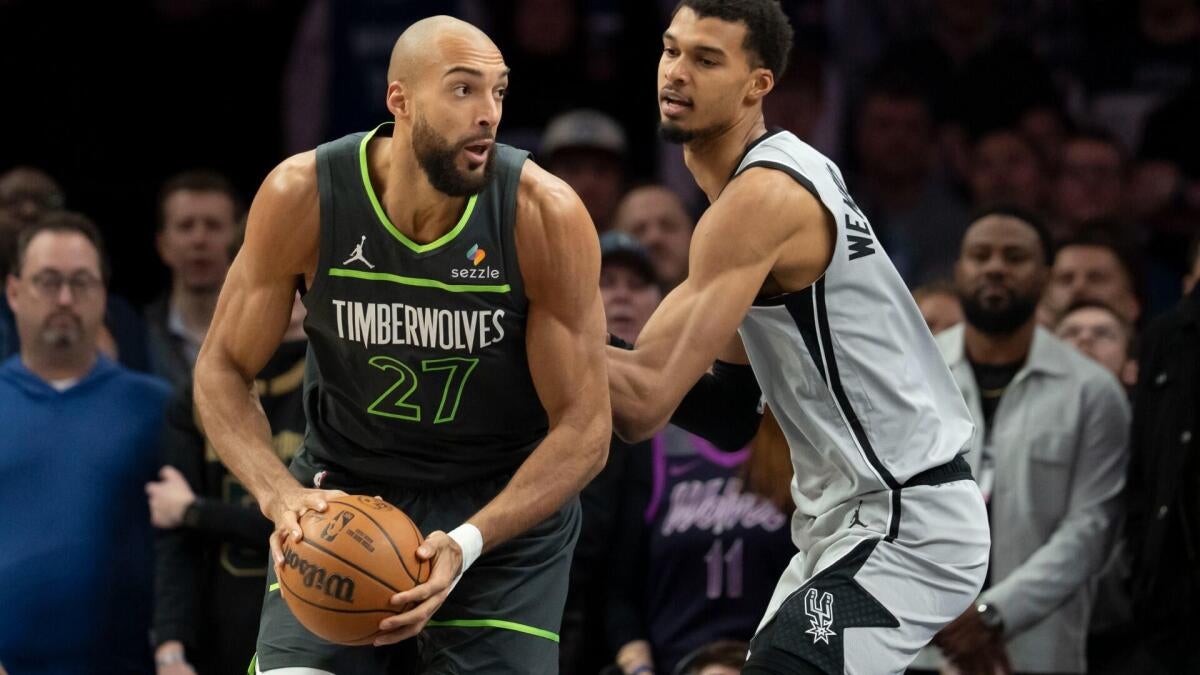

May 8, 2026, delivers a high-stakes slate of NBA Conference Semifinals and NHL Second Round showdowns. Betting markets…