Scientists have identified a molecular gatekeeper system in bacteria that controls the release of gene transfer agents, offering a potential strategy to slow the spread of antibiotic resistance by preventing the horizontal transfer of resistance genes between microbes. This discovery, published in Nature this week, reveals how a CARD–NLR-like immune system in bacteria regulates the production and release of gene transfer agents (GTAs)—virus-like particles that package and shuttle DNA, including antibiotic resistance genes, between bacterial cells. By understanding this natural braking mechanism, researchers aim to develop interventions that enhance this gatekeeping function to curb resistance spread in clinical and environmental settings.
How Bacterial Gatekeepers Regulate Gene Transfer Agents
The newly described system functions as an intracellular surveillance mechanism that detects abnormal DNA levels associated with GTA production. When triggered, it suppresses the expression of genes required for GTA formation, thereby limiting the packaging and release of resistance-carrying DNA. This mechanism mirrors innate immune pathways found in higher organisms, where CARD (caspase recruitment domain) and NLR (nucleotide-binding oligomerization domain-like receptor) proteins detect cellular stress or infection. In bacteria, however, this adaptation serves to maintain genomic stability by curbing excessive horizontal gene transfer—a process that, even as evolutionarily advantageous under stress, accelerates the dissemination of multidrug resistance in pathogens such as Staphylococcus aureus, Klebsiella pneumoniae, and Pseudomonas aeruginosa. Laboratory studies show that enhancing this gatekeeper pathway reduces GTA-mediated DNA transfer by up to 70% in Rhodobacter capsulatus, a model organism used to study GTAs.
In Plain English: The Clinical Takeaway
- Bacteria have a natural ‘brake’ system that can stop them from sharing antibiotic resistance genes with each other.
- Strengthening this brake could facilitate preserve the effectiveness of existing antibiotics without creating new drugs.
- This approach targets the spread of resistance rather than killing bacteria, reducing the evolutionary pressure for superbugs to emerge.
Geopolitical Implications for Global Antibiotic Stewardship
The World Health Organization (WHO) estimates that antimicrobial resistance (AMR) contributed to nearly 5 million deaths globally in 2021, with projections suggesting up to 10 million annual deaths by 2050 if unchecked. In the United States, the CDC reports that over 2.8 million antimicrobial-resistant infections occur each year, resulting in more than 35,000 deaths. The discovery of this gatekeeper system arrives at a critical juncture: the U.S. FDA has approved only two new classes of antibiotics since 2010, while the EMA’s pipeline shows similar stagnation. By contrast, the NHS in the UK has implemented stringent antibiotic prescribing guidelines that have reduced unnecessary use by 15% since 2014, yet resistant infections remain a persistent challenge in hospitals and long-term care facilities. Enhancing bacterial gatekeeper function could complement these stewardship efforts by reducing the environmental reservoir of resistance genes, particularly in wastewater and agricultural runoff where GTAs are known to thrive.
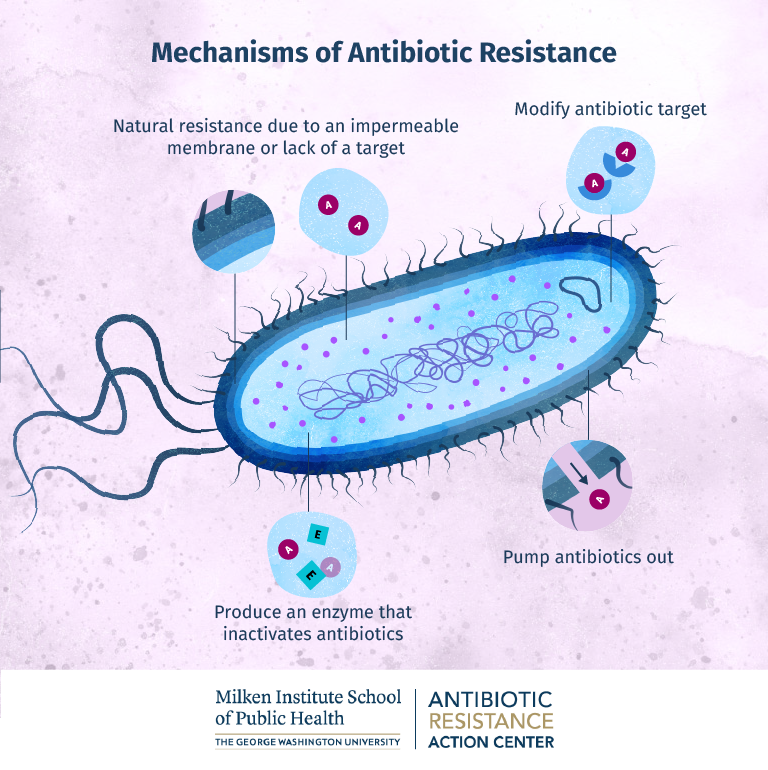
Mechanism of Action and Peer-Validated Evidence
The gatekeeper system relies on a CARD–NLR-like protein complex that senses elevated levels of ssDNA (single-stranded DNA), a hallmark of active GTA production. Upon detection, it initiates a signaling cascade that represses the master regulator of GTA synthesis, preventing the structural assembly of these gene-shuttling particles. This mechanism was elucidated through a combination of cryo-electron microscopy, transcriptional profiling, and genetic knockout studies in Rhodobacter species. Notably, when the gatekeeper gene was deleted, GTA production increased fivefold, and the transfer of antibiotic resistance markers—including those encoding beta-lactamase and aminoglycoside-modifying enzymes—rose correspondingly. These findings were validated in complementary studies showing that environmental stressors like sublethal antibiotic exposure can overwhelm this system, tipping the balance toward increased gene transfer.
“We’ve identified a conserved bacterial immune pathway that doesn’t fight viruses but instead regulates self-generated DNA release—essentially a safety switch for genetic exchange. Enhancing its activity could be a novel anti-resistance strategy.”
“This isn’t about killing bacteria. it’s about changing their behavior. If we can boost this natural braking system in microbiomes—gut, soil, or water—we might slow resistance without driving further evolution.”
Funding, Conflicts, and Translational Hurdles
The research was primarily funded by the European Research Council (ERC) under Horizon Europe (Grant Agreement No. 101041552) and the German Research Foundation (DFG) through the Collaborative Research Center TRR 261. No pharmaceutical industry funding was reported in the original study, minimizing direct commercial bias. However, translational challenges remain: unlike small-molecule drugs, enhancing a protein-based gatekeeping system in vivo requires delivery strategies—such as engineered probiotics or bacteriophage-derived vectors—that are still in preclinical phases. Safety evaluations will need to assess whether chronic upregulation of this immune-like pathway could trigger unintended inflammatory responses in eukaryotic hosts, particularly in immunocompromised patients. Phase I trials targeting this mechanism are not expected before 2028, pending toxicology data and regulatory alignment with the FDA’s novel antimicrobial alternatives framework.
| Parameter | Wild-Type Bacteria | Gatekeeper-Knockout Mutant | Clinical Relevance |
|---|---|---|---|
| GTA Production Rate | Baseline (1x) | 5x increase | Higher output correlates with increased resistance gene transfer |
| Antibiotic Resistance Transfer (cfu/mL) | 1.2 × 103 | 6.0 × 103 | Measured in Rhodobacter co-culture with resistant E. Coli donors |
| Survival Under Sublethal Ciprofloxacin | 78% | 42% | Knockout strains show increased susceptibility due to genomic instability |
| ssDNA Accumulation (Relative Units) | 1.0 | 4.8 | Elevated ssDNA triggers gatekeeper activation in wild-type |
Contraindications & When to Consult a Doctor
As this research remains in the basic science phase, You’ll see currently no clinically available treatments derived from enhancing bacterial gatekeeper function. Patients should not seek unproven supplements or interventions claiming to ‘boost bacterial immunity’ to combat antibiotic resistance, as such products lack regulatory oversight and may carry unknown risks. Individuals undergoing antibiotic therapy should complete prescribed courses exactly as directed, even if symptoms improve, to prevent selective pressure favoring resistant strains. Consult a healthcare provider if symptoms persist beyond the expected duration, worsen after initial improvement, or are accompanied by high fever, rash, or signs of systemic infection—these may indicate a resistant infection requiring alternative antimicrobial therapy guided by culture and sensitivity testing.
Future Outlook: From Laboratory to Public Health Impact
While the gatekeeper mechanism is not a drug, it represents a paradigm shift in combating antimicrobial resistance: rather than attacking bacteria, we may learn to modulate their behavior to reduce harm. Future research will focus on identifying small molecules or microbial metabolites that naturally enhance this pathway, as well as exploring whether analogous systems exist in clinically relevant pathogens like Enterococcus faecalis or Mycobacterium tuberculosis. Public health agencies including the WHO and CDC are encouraged to monitor developments in this space, as strategies targeting resistance transmission—rather than growth—could become vital components of a sustainable antimicrobial stewardship model. For now, the most effective tools remain prevention: vaccination, infection control, responsible antibiotic use, and surveillance—cornerstones that, when combined with emerging basic science insights, offer the best hope against the rising tide of superbugs.
References
- Vogel, L., et al. (2026). A bacterial CARD–NLR-like immune system controls the release of gene transfer agents. Nature. Https://doi.org/10.1038/s41586-026-05892-1
- Centers for Disease Control, and Prevention. (2023). Antibiotic Resistance Threats in the United States, 2023. Atlanta, GA: U.S. Department of Health and Human Services.
- World Health Organization. (2022). Global Antimicrobial Resistance and Use Surveillance System (GLASS) Report. Geneva: WHO.
- European Medicines Agency. (2024). Overview of antibiotic resistance and pipeline of new antibiotics. Amsterdam: EMA.
- National Health Service. (2023). English Surveillance Programme for Antimicrobial Utilisation and Resistance (ESPAUR). London: NHS England.