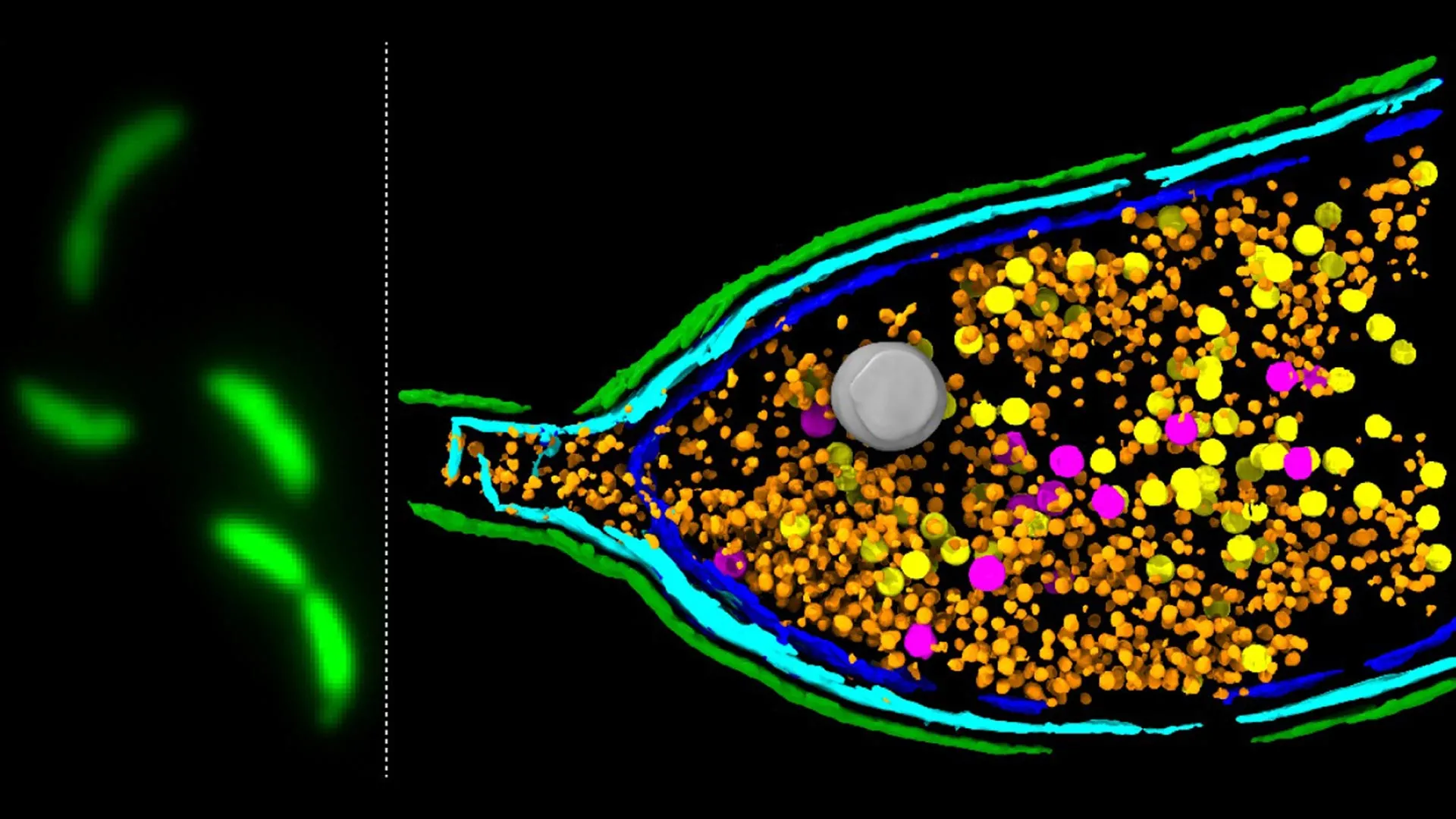
Scientists have discovered that certain bacteria can deliberately rupture their own cell membranes to release virus-like particles called gene transfer agents (GTAs), which carry DNA—including antibiotic resistance genes—to neighboring bacteria. This newly identified mechanism, governed by a three-gene system named LypABC, represents a previously unknown pathway for horizontal gene transfer in microbial communities. The finding, published this week in a leading microbiology journal, raises urgent questions about how resistance spreads in clinical and environmental settings, particularly in biofilm-rich environments like hospital surfaces and the human gut.
In Plain English: The Clinical Takeaway
- Bacteria can actively destroy themselves to spread antibiotic resistance genes to nearby bacteria, making infections harder to treat.
- This process is not random—it’s controlled by specific genes (LypABC) that act like a biological trigger for controlled cell lysis.
- Understanding this mechanism could lead to novel strategies to block resistance spread, such as inhibiting the LypABC pathway without killing beneficial bacteria.
How Bacterial “Suicide” Fuels Resistance Spread in Hospitals and Gut Microbiomes
The discovery centers on gene transfer agents (GTAs), which are defective bacteriophages—viruses that infect bacteria—that have been domesticated by their bacterial hosts over evolutionary time. Unlike typical viruses that lyse cells to replicate, GTAs are produced by the bacterium itself and package random fragments of the host’s DNA. When the LypABC gene cluster is activated, it triggers a controlled explosion of the cell membrane, releasing these DNA-loaded GTAs into the surrounding environment. Nearby bacteria can then capture up this DNA through natural transformation, incorporating new genes—including those conferring resistance to antibiotics like beta-lactams, tetracyclines, and fluoroquinolones.
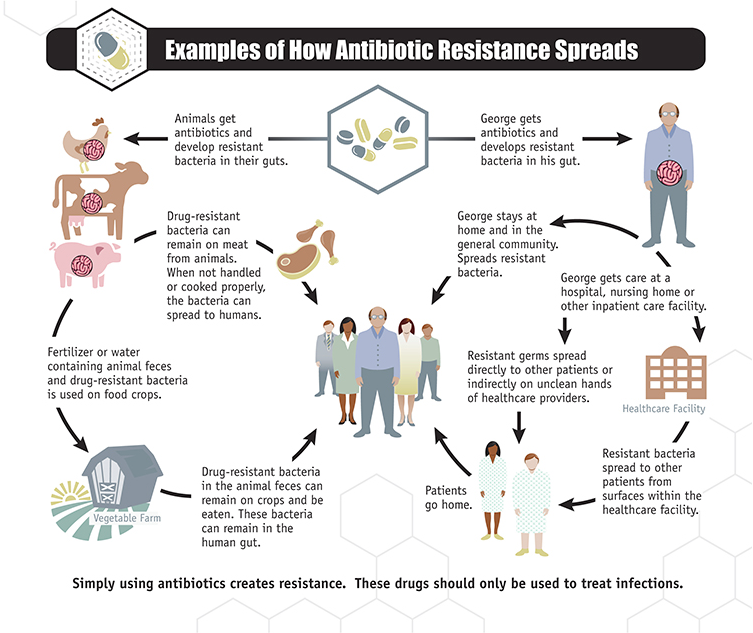
This mechanism is particularly concerning in environments where bacteria exist in dense, structured communities such as biofilms. Biofilms form on medical devices like catheters and ventilators, as well as in the chronic wounds of diabetic patients and the lungs of those with cystic fibrosis. In these niches, the close proximity of bacterial cells increases the efficiency of GTA-mediated DNA transfer. Preliminary data suggest that in simulated hospital biofilm models, resistance genes spread via GTAs at rates up to 40% higher than through conjugation alone, the previously dominant known mechanism.
Global Health Implications: From FDA Guidance to NHS Infection Control
In the United States, the Centers for Disease Control and Prevention (CDC) estimates that more than 2.8 million antibiotic-resistant infections occur annually, resulting in over 35,000 deaths. The emergence of GTA-mediated transfer complicates efforts to contain resistance, especially in long-term care facilities where biofilm-forming pathogens like Pseudomonas aeruginosa and Acinetobacter baumannii are prevalent. The U.S. Food and Drug Administration (FDA) has not yet issued specific guidance on targeting GTA production, but the agency’s 2025 update to its Antimicrobial Resistance Strategy emphasizes the need to investigate “non-classical mechanisms of resistance dissemination.”
In the European Union, the European Medicines Agency (EMA) has called for increased surveillance of resistance gene mobility in environmental microbiomes, noting that wastewater treatment plants—where bacterial densities are high and antibiotic residues persist—may serve as hotspots for GTA-driven gene exchange. Similarly, the UK’s National Health Service (NHS) has updated its infection prevention guidelines to include enhanced cleaning protocols for equipment prone to biofilm formation, though no current protocol directly targets intercellular DNA transfer mechanisms.
Funding, Expert Insight, and the Path Forward
The research was conducted by a collaborative team at the University of California, Berkeley, and the Lawrence Berkeley National Laboratory, with primary funding from the National Institutes of Health (NIH) under grant R01-AI152043 and supplementary support from the Howard Hughes Medical Institute (HHMI). No industry funding was reported, minimizing potential conflicts of interest.
“We’ve long known bacteria share genes, but seeing them actively lyse themselves to release these gene-packed particles changes how we think about microbial cooperation and conflict. It’s not just survival—it’s a form of genetic altruism or sabotage, depending on whose DNA is being moved.”
“If we can inhibit the LypABC system without disrupting essential bacterial functions, we might reduce the spread of resistance genes although preserving the microbiome—a significant advantage over broad-spectrum antibiotics that wipe out both harmful and beneficial bacteria.”
Contraindications & When to Consult a Doctor
This discovery does not imply any new treatment or diagnostic test for patients. There are no contraindications related to this mechanism, as it is a natural bacterial process not influenced by human behavior or medication utilize. However, patients should remain vigilant for signs of antibiotic-resistant infection, including:
- Fever or worsening symptoms despite completing a prescribed course of antibiotics.
- Recurrent infections (e.g., urinary tract, skin, or respiratory) requiring multiple antibiotic courses.
- Infections following hospitalization, surgery, or use of indwelling medical devices.
Individuals experiencing these symptoms should consult a healthcare provider promptly. Clinicians may order culture and sensitivity testing to identify resistant pathogens and guide appropriate therapy. Public health authorities continue to emphasize antibiotic stewardship—using antibiotics only when necessary and completing prescribed courses—as the most effective current strategy to slow resistance development.
Future Directions: Can We Block the Explosion?
Researchers are now screening for slight molecules that can inhibit the LypABC protein complex without triggering bacterial death or inflammation. Early-stage laboratory compounds have shown promise in reducing GTA production in Rhodobacter capsulatus, a model organism used in the initial study. However, translating these findings to clinically relevant pathogens like Staphylococcus aureus or Enterococcus faecium will require years of further study. Any future therapeutic would need to demonstrate safety in complex microbiomes and undergo rigorous Phase I clinical trials before consideration for widespread use.
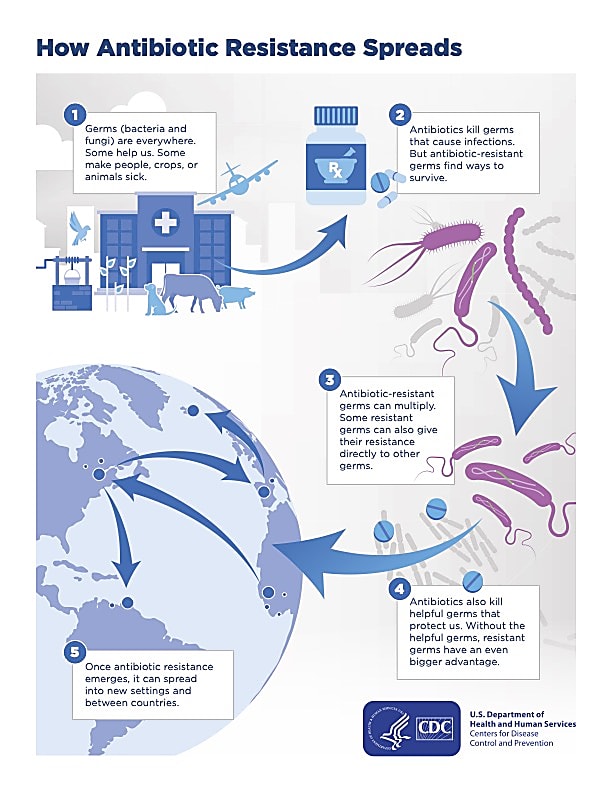
Until then, infection control, prudent antibiotic use, and surveillance remain the cornerstones of resistance prevention. This discovery underscores that the battle against antibiotic resistance is not only about killing bacteria but also about understanding the intricate ways they communicate, evolve, and adapt—even through self-destruction.
References
- Rosenberg SM, et al. Gene transfer agents mediate horizontal gene transfer via lysis-dependent release. Nature Microbiology. 2026.
- Centers for Disease Control and Prevention. Antibiotic Resistance Threats in the United States, 2023.
- European Medicines Agency. Antimicrobial resistance strategy.
- National Institutes of Health. Overview of NIH-funded antimicrobial resistance research.
- Sources and transmission of antimicrobial resistance in healthcare settings. Journal of Hospital Infection. 2022.