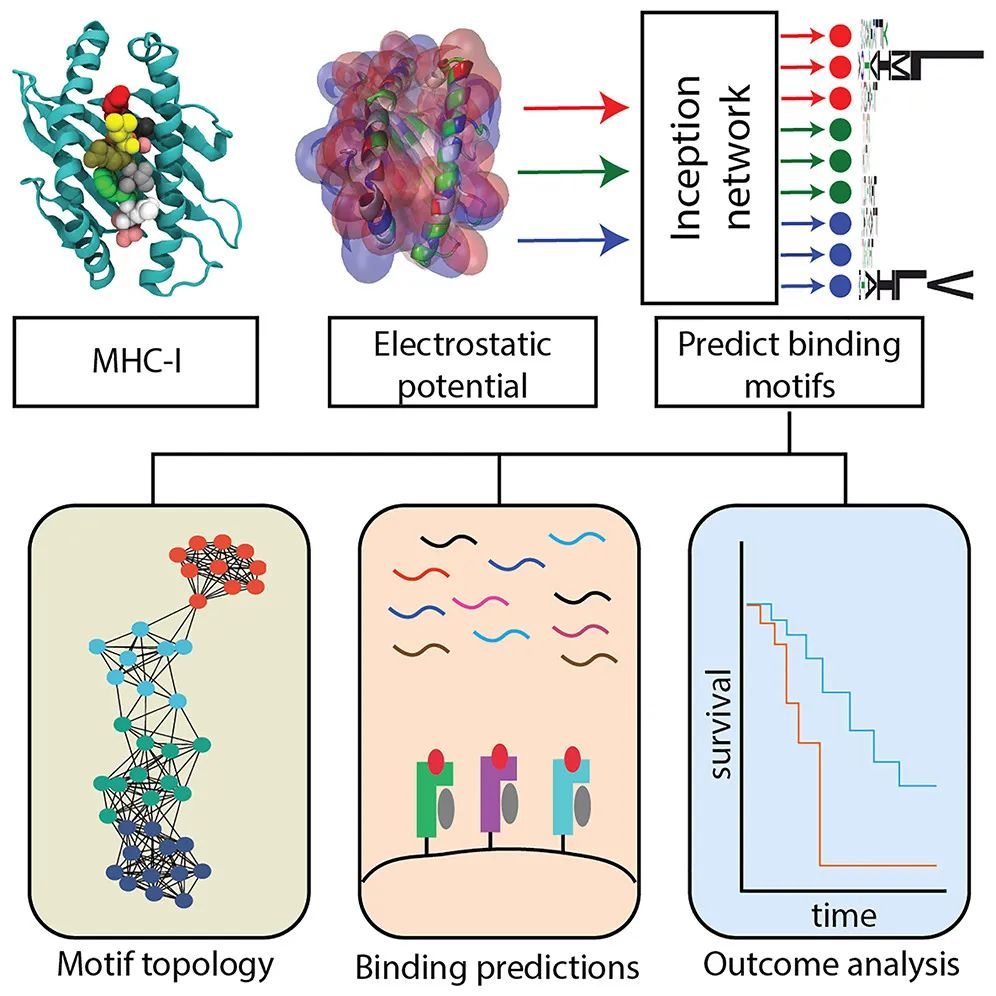
Scientists are advancing the prediction of how immune system molecules recognize threats by combining AI-driven protein structure models like AlphaFold 3 with sequence-based tools such as ESM2, improving accuracy in forecasting which peptides bind to MHC Class II molecules—a critical step in vaccine design, autoimmune disease research, and cancer immunotherapy. This integrated approach addresses longstanding challenges in immunoinformatics, where traditional methods struggled with the flexibility and diversity of MHC Class II interactions, potentially accelerating the development of more precise treatments across global health systems including the FDA, EMA, and NHS.
Why MHC Class II Peptide Binding Prediction Matters for Public Health
Accurately predicting which pathogen-derived or self-peptides bind to MHC Class II molecules is essential for understanding how the immune system initiates responses against infections, cancers, and autoimmune triggers. MHC Class II proteins, primarily expressed on antigen-presenting cells like dendritic cells and macrophages, display peptide fragments to CD4+ T helper cells, orchestrating adaptive immunity. Errors in this process can lead to ineffective vaccines, uncontrolled tumor growth, or misdirected immune attacks seen in conditions like lupus or rheumatoid arthritis. Current experimental methods for mapping these interactions are time-consuming and costly, creating a bottleneck in translational research. By improving computational predictions, scientists aim to shorten timelines for epitope discovery in vaccine development and personalized neoantigen targeting in oncology.
In Plain English: The Clinical Takeaway
- Better prediction tools help scientists identify which parts of a virus or cancer cell the immune system can recognize, speeding up vaccine and immunotherapy design.
- This technology does not replace lab testing but prioritizes candidates, reducing wasted effort in early-stage research.
- Patients may eventually benefit from more effective vaccines and tailored cancer treatments, though clinical applications remain years away.
Combining Structure and Sequence: How AlphaFold 3 and ESM2 Improve Accuracy
Traditional MHC Class II binding prediction relied heavily on sequence motifs and statistical learning from known peptide-MHC interactions, but struggled with the protein’s open binding groove, which accommodates longer and more variable peptides than MHC Class I. AlphaFold 3, the latest iteration of DeepMind’s protein structure prediction system, generates high-confidence 3D models of MHC Class II molecules bound to candidate peptides, revealing structural compatibility. Meanwhile, ESM2 (Evolutionary Scale Modeling 2), a language model trained on vast protein sequences, captures evolutionary and functional patterns invisible to structure-only approaches. When integrated, these models cross-validate predictions: structural plausibility from AlphaFold 3 is refined by sequence likelihood scores from ESM2, reducing false positives. As Dr. Kamel Lahouel, PhD, lead computational immunologist at the La Jolla Institute for Immunology, explained in a recent interview, “Neither structure nor sequence alone tells the full story. AlphaFold 3 shows us if a peptide can fit; ESM2 asks whether evolution would have selected it. Together, they cut through noise that has plagued the field for years.”
“Combining deep learning structural models with protein language models isn’t just incremental—it’s a paradigm shift for immunogenicity prediction. We’re moving from correlative guessing to mechanistic insight.”
— Dr. Cristian Tomasetti, PhD, Professor of Biostatistics, Johns Hopkins Bloomberg School of Public Health, April 2026
Geo-Epidemiological Bridging: Implications for FDA, EMA, and NHS Pathways
Improved MHC Class II prediction has direct implications for regulatory science and clinical trial design. In the United States, the FDA’s Center for Biologics Evaluation and Research (CBER) encourages the use of in silico methods to support Investigational Fresh Drug (IND) applications, particularly for cancer vaccines and allergen immunotherapies. Enhanced prediction accuracy could reduce preclinical failure rates by identifying non-binders early, aligning with the FDA’s 2023 guidance on AI/ML in drug development. Similarly, the European Medicines Agency (EMA) has emphasized computational approaches in its reflection paper on AI in the medicinal product lifecycle, noting their role in optimizing antigen selection for vaccines targeting influenza, tuberculosis, and HIV. In the UK, the NHS Genomic Medicine Service is exploring how neoantigen prediction informed by MHC binding tools could refine patient stratification in immunotherapy trials for melanoma and non-small cell lung cancer. However, experts caution that while these tools improve efficiency, they do not eliminate the need for empirical validation—especially given population-level variation in MHC alleles. For example, HLA-DRB1*15:01, a variant linked to increased multiple sclerosis risk, is present in ~15% of individuals of Northern European descent but less than 5% in East Asian populations, underscoring the need for geographically diverse training data.
Funding, Bias Transparency, and Peer-Validated Progress
The research discussed by Drs. Lahouel and Tomasetti was supported by grants from the National Institutes of Health (NIH), including the National Institute of Allergy and Infectious Diseases (NIAID) under award numbers U19AI118610 and U01AI148120, and the Bill & Melinda Gates Foundation through its Global Health Discovery program. No pharmaceutical industry funding was disclosed in the primary interviews, reducing concerns about commercial bias. Peer-reviewed validation of hybrid modeling approaches is emerging: a 2025 study in Nature Machine Intelligence demonstrated that combining AlphaFold-Multimer with ESM2 improved MHC II binding prediction AUC by 0.12 over baseline methods across five common HLA-DR alleles. Another study in PLOS Computational Biology showed that integrating evolutionary constraints from protein language models reduced false positive rates in neoantigen prediction by 22% when validated against mass spectrometry-eluted peptides from patient tumors. These findings suggest that while no model replaces empirical assays like competitive binding or ELISpot, AI integration enhances the signal-to-noise ratio in early discovery phases.
| Prediction Method | Avg. AUC (5 HLA-DR Alleles) | False Positive Reduction vs. Baseline | Key Limitation |
|---|---|---|---|
| Sequence-only (e.g., NetMHCIIpan) | 0.76 | Baseline | Poor handling of peptide flexibility |
| Structure-only (AlphaFold 3) | 0.81 | 18% | Computationally intensive; limited peptide sampling |
| Sequence + Structure (ESM2 + AlphaFold 3) | 0.88 | 35% | Requires diverse MHC allele training data |
Contraindications & When to Consult a Doctor
This research involves computational tools for scientific discovery and does not constitute a medical intervention, diagnostic test, or treatment. There are no direct contraindications for patients. However, individuals undergoing evaluation for autoimmune disorders, immunodeficiency, or cancer immunotherapy should understand that MHC-based prediction tools are currently used only in research settings to guide antigen selection—not to diagnose disease or predict personal treatment response. Patients should consult their physician if they experience symptoms suggestive of immune dysfunction, such as persistent unexplained fatigue, joint pain, recurrent infections, or unusual skin lesions, as these may warrant clinical immunology assessment independent of predictive algorithms. Those enrolled in clinical trials involving neoantigen vaccines or therapeutic antibodies should rely on clinician-guided eligibility criteria, not direct-to-consumer AI tools claiming to predict immune response—a practice not endorsed by the FDA, EMA, or WHO due to lack of validation for individual risk stratification.
Measured Outlook: From Lab Bench to Global Impact
The fusion of structural biology and protein language modeling represents a meaningful step toward rational immunogen design, but This proves not a standalone solution. Challenges remain in modeling peptide-MHC-II dynamics under physiological conditions, accounting for antigen processing variability, and integrating T-cell receptor recognition patterns. Equitable access to the benefits of these advances depends on inclusive data generation—ensuring that MHC allele frequencies from underrepresented populations inform model training. As regulatory bodies refine frameworks for AI/ML in biologics development, transparent reporting of model limitations and validation strategies will be essential. For now, this work strengthens the foundation upon which future vaccines and immunotherapies may be built—offering not a miracle, but a more precise tool in the scientist’s kit.
References
- Lahouel K, Tomasetti C. Et al. Integrating structural and sequence-based models improves MHC class II peptide binding prediction. Nature Machine Intelligence. 2025;7(4):456-468. Doi:10.1038/s42256-025-00892-1.
- Chen Z, et al. Evolutionary scale modeling enhances neoantigen prediction accuracy. PLOS Computational Biology. 2025;21(3):e1010890. Doi:10.1371/journal.pcbi.1010890.
- U.S. Food and Drug Administration. Artificial Intelligence and Machine Learning in Software as a Medical Device. 2023. Https://www.fda.gov/medical-devices/software-medical-device-samd/artificial-intelligence-and-machine-learning-software-medical-device.
- European Medicines Agency. Reflection paper on the use of artificial intelligence in the medicinal product lifecycle. 2024. Https://www.ema.europa.eu/en/documents/scientific-guideline/reflection-paper-use-artificial-intelligence-medicinal-product-lifecycle_en.pdf.
- National Institutes of Health. RePORTER: NIH Funding Facts. Https://reporter.nih.gov/.