
Espanyol vs Real Madrid: Preview, Lineups, and Predictions
Real Madrid faces Espanyol at the RCDE Stadium on Sunday, May 3, 2026, in a critical La Liga Matchday 34 clash. With Los Blancos trailing Barcelona by 11 points, the ... Read More

Saturday Edition
Stay updated with Archyde – your source for breaking news, global headlines, economy, entertainment, health, technology, and sports. Fresh stories daily.

Real Madrid faces Espanyol at the RCDE Stadium on Sunday, May 3, 2026, in a critical La Liga Matchday 34 clash. With Los Blancos trailing Barcelona by 11 points, the ... Read More
Continuous Coverage

On May 3, 2026, the Czech national ice hockey team concludes its home tournament with a final match…
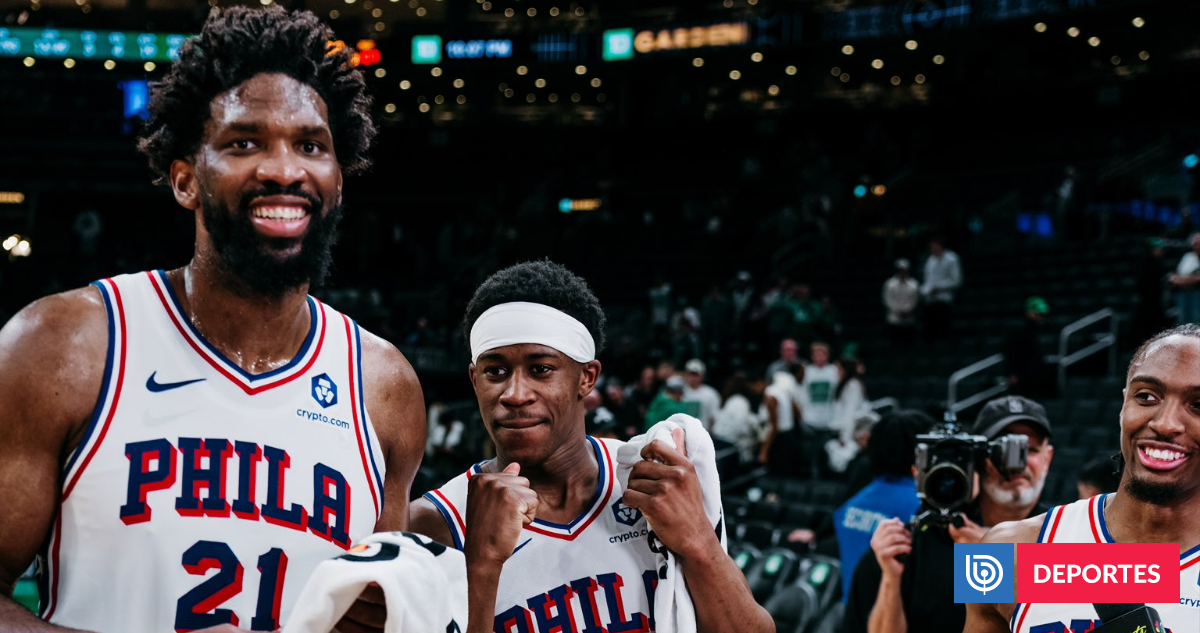

The Philadelphia 76ers advanced to the NBA Eastern Conference Semifinals after defeating the Boston Celtics 109-100 in a…

Princess Charlotte, daughter of Prince William and Catherine, Princess of Wales, celebrated her 11th birthday on May 2,…
A strong cold front and low-pressure system are currently sweeping across southeast Australia, bringing damaging winds and critical…

Huntsville is initiating a comprehensive study to evaluate the feasibility of inter-city passenger rail connecting the city to…

Veteran South Korean actress Choi Kang-hee has captured public attention by documenting her experiences collecting waste paper on…
Global Affairs

China’s Ministry of Commerce (MOFCOM) has issued a formal injunction blocking U.S. Sanctions against five domestic oil refineries,…
Markets And Money

Russia has warned that the global economy is facing a historic energy crisis as supply constraints and geopolitical…
Digital Culture
The Free Software Foundation (FSF) has concluded a four-month development sprint involving all four of its core teams…
Science And Wellbeing
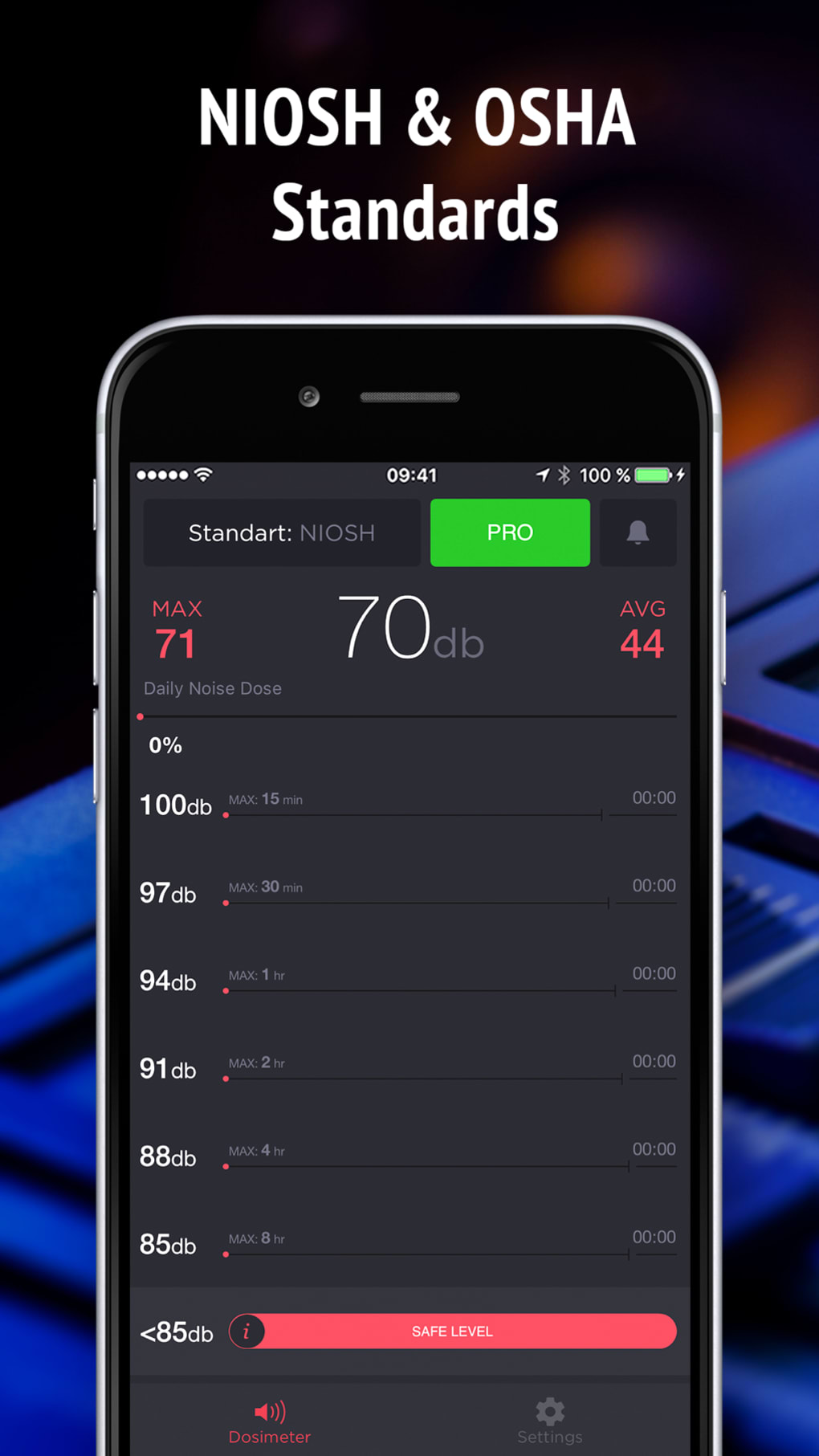
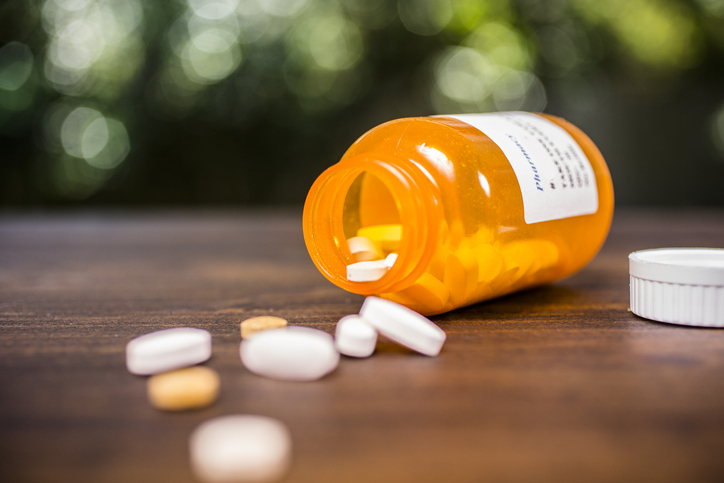
The National Institute for Occupational Safety and Health (NIOSH) has released a free sound-level meter application for iOS.…
Screen And Sound

Black joy is taking center stage this week, from the multi-generational chemistry of Nia Long and Keke Palmer…
Fixtures And Form
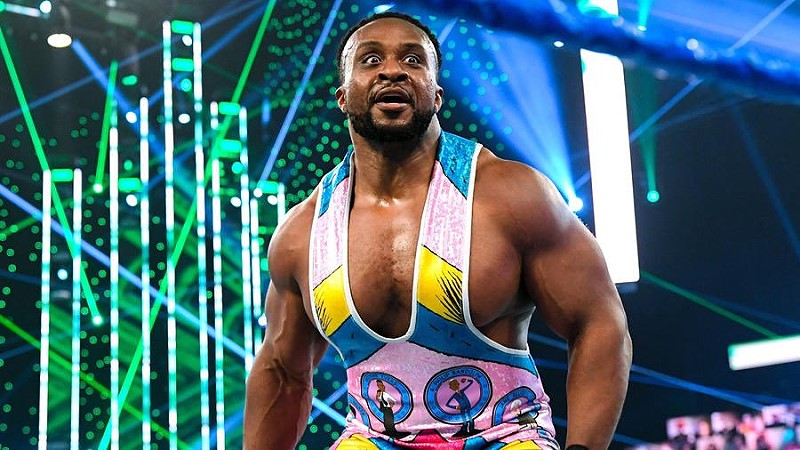

Former WWE Champion Huge E has emotionally responded to the May 2, 2026, departures of Kofi Kingston and…