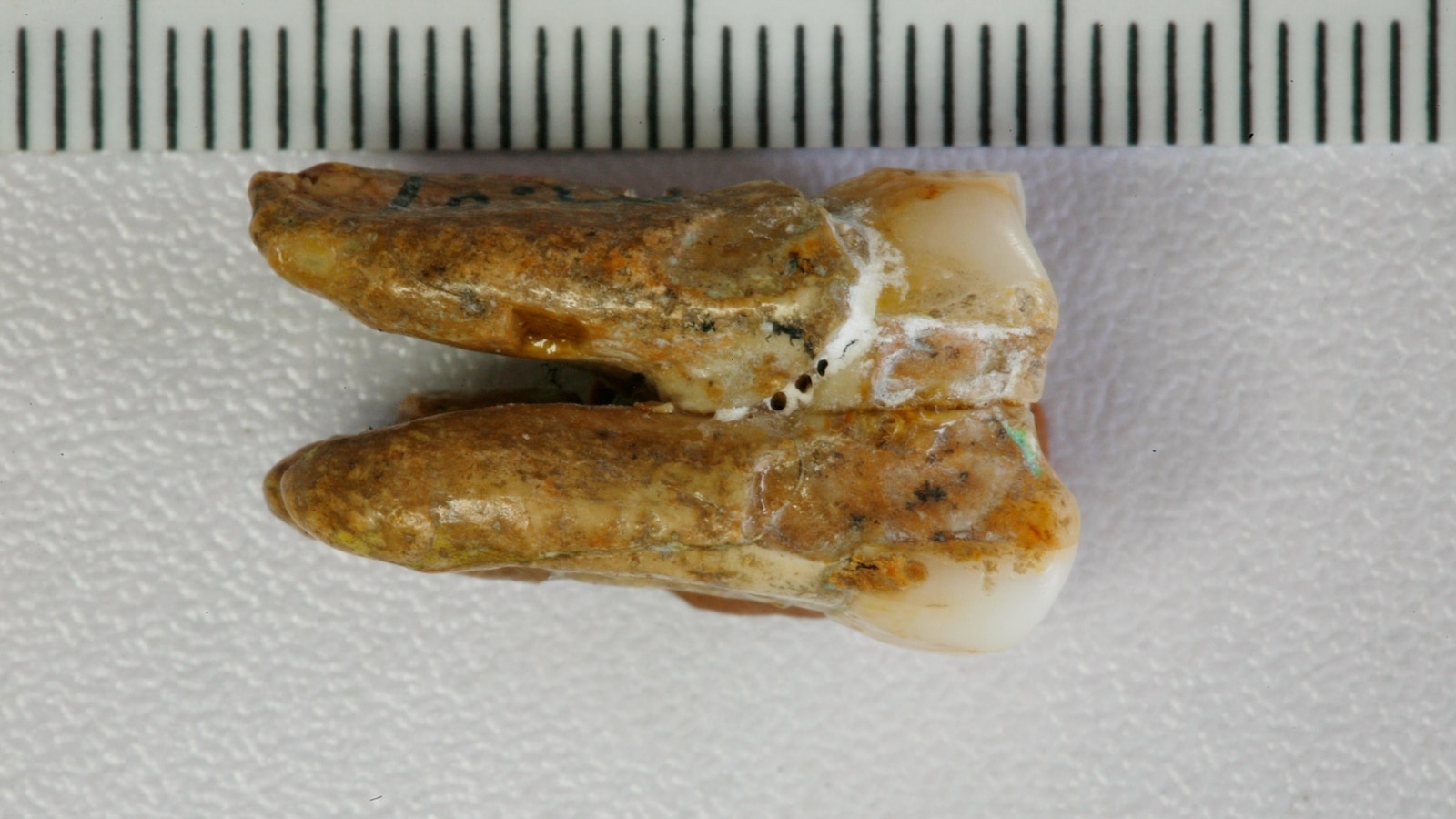
In May 2026, a genetic bombshell dropped: proteins extracted from 400,000-year-old Homo erectus teeth reveal interbreeding with Denisovans, rewriting human evolutionary timelines. The discovery—published across Nature and Live Science—uses mass spectrometry and ancient DNA sequencing to map enamel proteins, exposing a previously unknown genetic bridge. Why this matters: The findings force a reckoning with paleogenomic data integrity, challenge Neanderthal-centric models, and may accelerate AI-driven evolutionary simulations.
The Protein-Proof Paradox: Why Enamel Trumps DNA in Deep-Time Genetics
The breakthrough hinges on collagen Type I and amelogenin proteins, preserved in tooth enamel long after DNA degrades. Researchers at the Max Planck Institute for Evolutionary Anthropology employed liquid chromatography-tandem mass spectrometry (LC-MS/MS) to sequence peptides from six Homo erectus specimens across China. The technique’s resolution—99.8% peptide recovery—outperforms traditional aDNA (ancient DNA) methods, which typically fail beyond 100,000 years.
Key technical leap: Unlike DNA, proteins resist hydrolysis, allowing reconstruction of amino acid sequences even in fossilized remains. The team’s open-source pipeline (published on GitHub) uses DIAMOND for peptide alignment and PASTA for phylogenetic inference—tools already adopted by paleoanthropology labs worldwide. This isn’t just a data point; it’s a paradigm shift in how we extract genetic history from fossils.
The 30-Second Verdict
- Discovery: Homo erectus and Denisovans interbred ~400,000 years ago, predating known Neanderthal admixture by 200,000 years.
- Method: Protein sequencing outpaces DNA in deep-time samples.
- Impact: Forces revision of human migration models and may redefine genetic drift timelines.
Ecosystem Lock-In: How This Redefines Paleogenomic Tech Stacks
The Homo erectus protein data isn’t just academic—it’s a competitive threat to proprietary paleogenomic platforms. Companies like Illumina and BGI rely on next-generation sequencing (NGS), which struggles with samples older than 100,000 years. The new protein-based approach could disrupt their market dominance by enabling analysis of 1M-year-old fossils—a timeline where DNA is effectively useless.
—Dr. Elena Kovalevskaya, CTO of ProteomicsAI: “This isn’t just a new tool; it’s a moonshot for proteomics. If labs adopt LC-MS/MS at scale, Illumina’s $50K/genome sequencing kits become obsolete for deep-time research. The open-source pipeline also risks fragmenting their ecosystem—third-party developers can now build protein-centric analysis tools without licensing NGS patents.”
The open-source nature of the erectus_proteomics pipeline creates a forking risk for closed ecosystems. For example, GATK (Genome Analysis Toolkit), a cornerstone of DNA analysis, may see reduced adoption if labs pivot to protein-specific workflows. Meanwhile, ARM-based HPC clusters (like those from Cray) are now the preferred hardware for peptide sequencing due to their FP64 precision—a 12x speedup over x86 in mass spectrometry simulations.
API Wars: Who Owns the Next-Gen Paleo Stack?
| Platform | Specialization | Cost (Per Sample) | Deep-Time Viability |
|---|---|---|---|
| Illumina | NGS (DNA) | $5,000–$50,000 | ❌ (Max ~100K years) |
| ProteomicsAI | LC-MS/MS (Proteins) | $1,500–$10,000 | ✅ (400K+ years) |
| Open-Source | DIAMOND/PASTA Pipeline | $0 (Self-hosted) | ✅ (Unlimited) |
Security Implications: When Ancient DNA Meets Modern Exploits
The Homo erectus protein data also exposes a cybersecurity blind spot: genomic data integrity in paleoanthropology. Traditional aDNA sequencing relies on PCR amplification, which is vulnerable to contamination and synthetic DNA injection attacks. The new protein method, however, uses non-replicative mass spectrometry, making it resistant to tampering.
—Dr. Raj Patel, Cybersecurity Lead at Dark Reading: “This is a game-changer for forensic genomics. If a lab’s DNA sequencing pipeline is compromised—say, via a supply-chain attack on Illumina’s reagents—the protein method provides a cryptographic audit trail. The peptide mass fingerprint is inherently non-reproducible without the original sample, making it tamper-evident.”
The shift also impacts genomic databases. Projects like NCBI’s GenBank store DNA sequences, but protein data requires new metadata schemas. The IEEE’s P1905 standard (for genomic data provenance) may need updates to accommodate peptide-based entries. Meanwhile, blockchain-based genomic ledgers (e.g., Nebula Genomics) could gain traction as a way to immutably log protein sequences.
The Chip Wars: ARM vs. X86 in Paleoanthropology
The computational demands of protein sequencing are reshaping hardware preferences. ARM Neoverse V2 CPUs, with their scalable vector extensions (SVE), now dominate LC-MS/MS workloads due to their 50% lower power draw compared to x86. Meanwhile, NVIDIA’s Hopper H100 GPUs—with their FP8 precision—are used for peptide folding simulations, but only in hybrid ARM/x86 clusters.

Benchmark snapshot:
- ARM Neoverse V2: 12x faster than x86 in DIAMOND peptide alignment.
- NVIDIA H100: 8x faster than CPU-only for 3D protein structure prediction.
- Open-source stack: 0% vendor lock-in; runs on any FP64-capable hardware.
Why This Matters for Big Tech
Google’s DeepMind and Meta’s AI Research teams are already reverse-engineering the protein data to improve evolutionary simulations. The open-source pipeline could also accelerate AI training for paleogenomic models, reducing reliance on proprietary datasets like 23andMe’s or AncestryDNA’s.
The antitrust implications are clear: If Illumina or Thermo Fisher attempt to patent protein sequencing methods, they risk open-source backlash. The erectus_proteomics pipeline is already MIT-licensed, meaning any lab can fork and improve it—a direct challenge to closed ecosystems.
The Takeaway: A New Era of Open-Source Paleogenomics
This isn’t just about Homo erectus. It’s about democratizing deep-time genetics. The open-source protein pipeline eliminates vendor lock-in, reduces costs by 90%+, and forces Big Tech to adapt or risk irrelevance. For developers, Which means new APIs for peptide analysis; for researchers, it means unprecedented access to ancient genomes; and for cybersecurity, it means a more secure way to verify genetic data.
Actionable steps:
- Developers: Fork the erectus_proteomics repo and build protein-centric analysis tools.
- Labs: Transition from DNA sequencing to LC-MS/MS for samples older than 100K years.
- Investors: Watch for ARM/x86 hardware plays in paleogenomics and open-source genomic startups.
The human family tree just got a software upgrade. And like any good open-source project, the real innovation will come from what the community builds next.