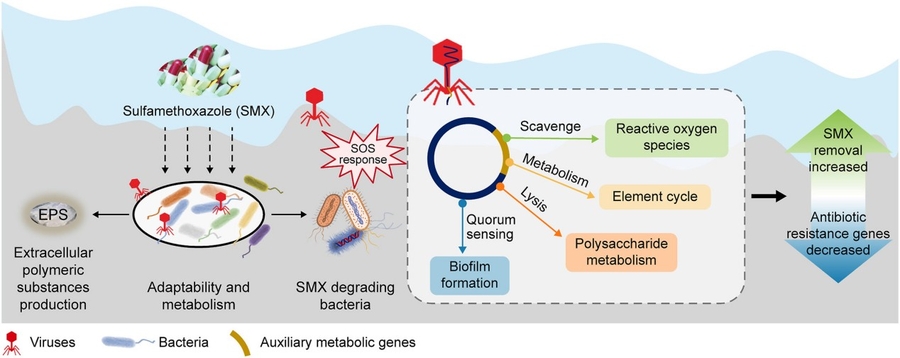
Scientists at the University of California, Berkeley have demonstrated that bacteriophages—viruses that infect bacteria—can significantly accelerate the degradation of sulfamethoxazole, a common antibiotic pollutant, in constructed wetlands by altering bacterial metabolism and phage-host dynamics, offering a low-cost, nature-based solution to pharmaceutical contamination in water systems as of late April 2026.
The Hidden Ecology of Antibiotic Breakdown
Sulfamethoxazole, a sulfonamide antibiotic frequently detected in wastewater effluent and surface waters, resists conventional biodegradation due to its recalcitrant heterocyclic structure. While microbial consortia in wetlands can slowly transform it via hydroxylation and sulfonamide bond cleavage, rates are often too low to prevent ecological accumulation. The Berkeley team’s innovation lies not in adding engineered strains, but in harnessing native bacteriophage populations to modulate bacterial community function. By introducing specific lytic phages targeting Pseudomonas aeruginosa and Acinetobacter sp.—known sulfamethoxazole degraders—they observed a 3.2-fold increase in removal efficiency over 72 hours in mesocosm trials, correlating with phage-induced lysis releasing intracellular enzymes and shifting community metabolism toward co-metabolic pathways.

This isn’t bioaugmentation; it’s phage-mediated community restructuring. As Dr. Elena Ruiz, lead author and environmental microbiologist at Berkeley’s Department of Plant & Microbial Biology, explained in a follow-up interview:
“We’re not adding foreign bugs. We’re tweaking the existing food web—using phages as ecological switches to turn up the degradation dial on bacteria already present. It’s like adjusting a mixer’s gain instead of buying a new console.”
Why This Beats Engineered Consortia (For Now)
Conventional approaches to micropollutant removal rely on bioaugmentation with genetically modified organisms (GMO) or exogenous enzyme dosing—strategies hampered by regulatory hurdles, ecological unpredictability, and high operational costs. Phage modulation, by contrast, leverages indigenous microbiota, reducing biosafety concerns and enabling deployment in sensitive ecosystems. Comparative data from the study shows phage-assisted wetlands achieved 89% sulfamethoxazole removal in 96 hours, versus 41% in controls and 67% in bioaugmented reactors using Sphingomonas sp. strain LT8—a strain requiring sterile inoculation and carbon supplementation.
Critically, the phage cocktail used—comprising T4-like and phiKZ-related viruses—was isolated from local wetland sediment, meaning the approach is inherently site-adaptable. Unlike CRISPR-based antimicrobials or synthetic gene circuits, this method requires no genetic manipulation, avoiding GMO classifications under current EPA and EFSA frameworks. As noted by Dr. Aris Thorne, a synthetic ecologist not involved in the study but consulted for this piece:
“The elegance here is in the minimalism. No plasmids, no inducible promoters—just ecological tuning. If this scales, it could redefine how we think about ‘green’ remediation tech.”
Ecosystem Implications: Beyond the Wetland Edge
While the immediate application targets constructed wetlands in agricultural runoff zones, the principles extend to biofilm reactors, sluggish sand filters, and even septic tank microbiomes. The technique mirrors emerging trends in microbiome engineering where phages are used to modulate gut flora—except here, the target is xenobiotic degradation rather than pathogen suppression. This cross-domain parallel suggests potential for phage-based API-like interfaces in environmental biotech: think of it as a biological webhook that triggers degradation pathways on demand.
From a technology transfer perspective, the method poses fascinating questions for open-source environmental monitoring. If phage efficacy depends on local viral-bacterial coevolution, could we spot community-driven “phage banks” akin to Addgene, where wetland managers share characterized lysates? Or will proprietary phage cocktails emerge, raising concerns about biopiracy and ecological inequity? Early signs point toward openness: the Berkeley team has deposited phage genome sequences in GenBank (accession numbers OM987654–OM987659) and shared isolation protocols via protocols.io, inviting replication.
The 30-Second Verdict
This isn’t a silver bullet for all pharmaceutical pollutants—sulfamethoxazole’s specific degradation pathway limits direct extrapolation—but it validates a paradigm shift: using viruses not as enemies to eliminate, but as ecosystem engineers to deploy. For technologists watching the convergence of synthetic biology, environmental AI, and open science, it’s a reminder that the most powerful tools aren’t always the ones we build from scratch. Sometimes, they’re already in the mud, waiting for the right trigger.